Mastering NIR Photon Migration: A Complete Guide to Monte Carlo Simulation for Biomedical Optics
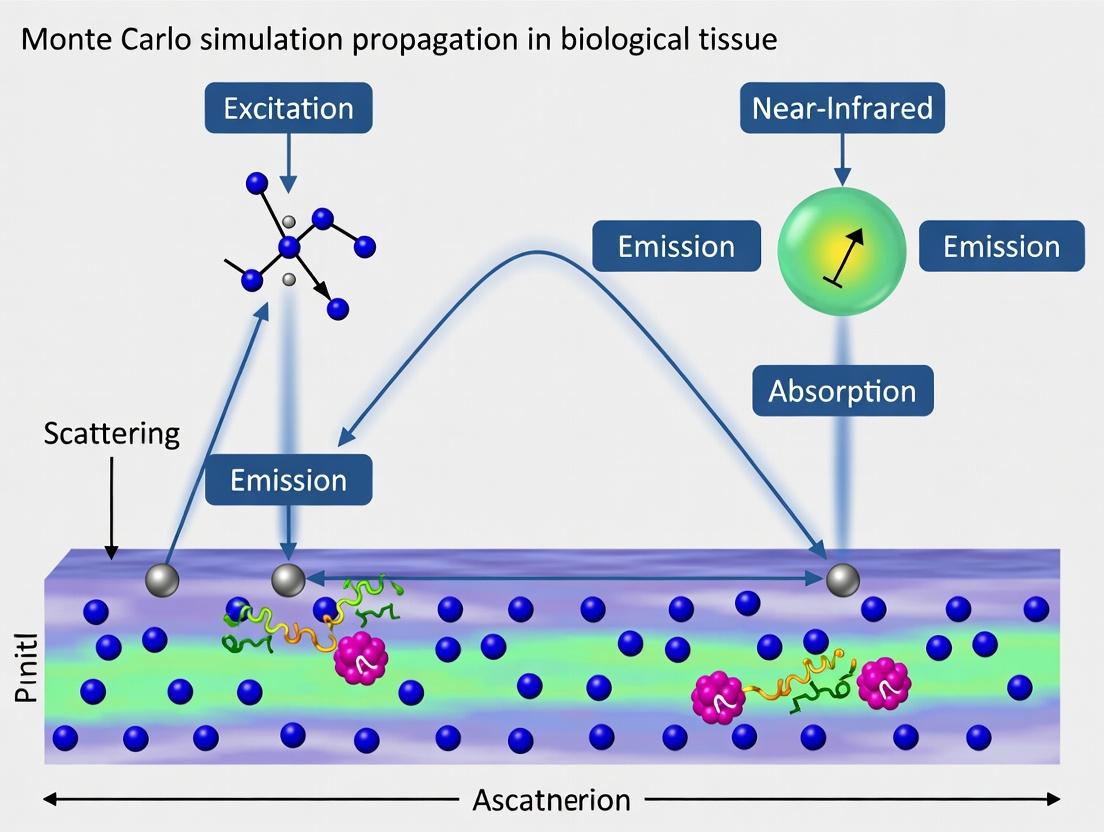
This article provides a comprehensive guide to Monte Carlo simulations for modeling near-infrared (NIR) photon propagation in biological tissues.
Mastering NIR Photon Migration: A Complete Guide to Monte Carlo Simulation for Biomedical Optics
Abstract
This article provides a comprehensive guide to Monte Carlo simulations for modeling near-infrared (NIR) photon propagation in biological tissues. We first establish the fundamental principles of photon-tissue interactions and the rationale for stochastic modeling. We then detail the step-by-step methodology for building accurate simulations, from code architecture to modeling complex tissue geometries, with direct applications in diffuse optical imaging and spectroscopy. The guide addresses common challenges, including computational bottlenecks and source/detector modeling, offering practical troubleshooting and optimization strategies. Finally, we cover critical validation techniques against analytical solutions and phantom studies, and compare major simulation platforms (e.g., MCX, TIM-OS, Monte Carlo eXtreme). Designed for researchers, scientists, and drug development professionals, this resource synthesizes current best practices to enable robust, efficient, and reliable simulations for advancing non-invasive diagnostics and therapeutic monitoring.
The Physics of Light in Tissue: Why Monte Carlo is Essential for Modeling NIR Photon Migration
This whitepaper provides a technical foundation on photon-tissue interactions within the near-infrared (NIR) window, specifically scoped for Monte Carlo (MC) simulations of light propagation in biological tissues. We detail the core physical phenomena—scattering, absorption, and anisotropy—that govern the spatial and temporal distribution of photon energy, which is critical for modeling applications in biomedical optics, drug development, and therapeutic monitoring.
Core Principles of Photon-Tissue Interaction
Photon transport in tissue is governed by the radiative transfer equation (RTE). For computational tractability in complex geometries, the stochastic Monte Carlo method is the gold standard for solving the RTE. The fundamental interactions considered are absorption and scattering, characterized by the absorption coefficient (μa) and scattering coefficient (μs). A critical derived parameter is the reduced scattering coefficient (μs' = μs(1 - g)), which describes scattering in the diffusion-dominated regime.
The NIR Window
The "NIR window" or "therapeutic window" (approximately 650-1350 nm) is a spectral region where light penetration into biological tissue is maximized. This occurs due to relatively low absorption from endogenous chromophores like hemoglobin, water, and lipids, allowing photons to probe deep tissues.
Table 1: Optical Properties of Key Tissue Chromophores in the NIR Window
| Chromophore | Primary Absorption Peaks (nm) | Function in Photon Interaction | Typical μa in Tissue (650-900 nm) [cm⁻¹] |
|---|---|---|---|
| Oxyhemoglobin (HbO₂) | ~415, 542, 577, ~925 | Dominant absorber in blood; informs oximetry | 0.03 - 0.3 (varies with conc.) |
| Deoxyhemoglobin (Hb) | ~430, 555, ~760 | Key absorber; informs oxygen saturation | 0.02 - 0.25 (varies with conc.) |
| Water (H₂O) | ~980, >1150 | Becomes significant absorber >900 nm | ~0.02 (at 900 nm) to >1.0 (at 1150 nm) |
| Lipid | ~930, 1040, 1210 | Absorber in adipose tissue and cell membranes | ~0.03 - 0.1 (at 930 nm) |
Table 2: Typical Optical Properties of Biological Tissues in the NIR Window (at 800 nm)
| Tissue Type | Absorption Coefficient, μa (cm⁻¹) | Scattering Coefficient, μs (cm⁻¹) | Anisotropy Factor, g | Reduced Scattering Coefficient, μs' (cm⁻¹) |
|---|---|---|---|---|
| Skin (epidermis/dermis) | 0.1 - 0.2 | 150 - 200 | 0.7 - 0.9 | 15 - 45 |
| Brain (gray matter) | 0.15 - 0.35 | 150 - 250 | 0.8 - 0.95 | 10 - 40 |
| Breast Tissue | 0.03 - 0.08 | 100 - 150 | 0.7 - 0.95 | 5 - 30 |
| Skeletal Muscle | 0.1 - 0.3 | 200 - 300 | 0.8 - 0.95 | 10 - 50 |
| Adipose Tissue | 0.05 - 0.15 | 100 - 200 | 0.7 - 0.9 | 10 - 50 |
| Liver | 0.3 - 0.6 | 200 - 350 | 0.8 - 0.95 | 15 - 50 |
Scattering
Scattering in tissue is primarily elastic (Mie scattering) due to structures like organelles, cell membranes, and collagen fibers. It changes the photon's direction without energy loss. The scattering coefficient (μs) represents the probability of scattering per unit path length. The phase function, often approximated by the Henyey-Greenstein function, describes the angular distribution of scattered light.
Absorption
Absorption converts photon energy into other forms (e.g., heat, fluorescence). The absorption coefficient (μa) is the probability of absorption per unit path length. It depends on the concentration and absorption spectra of chromophores in the tissue.
Anisotropy
The anisotropy factor (g) is the average cosine of the scattering angle, ranging from -1 (perfect backscatter) to 1 (perfect forward scatter). Biological tissues are highly forward-scattering, with typical g values of 0.7-0.95 in the NIR. This parameter is crucial for correctly modeling photon paths in MC simulations.
Monte Carlo Simulation: A Modeling Framework
Monte Carlo methods track individual photon packets as they undergo random walks based on probability distributions derived from μa, μs, and g. The core algorithm involves:
- Photon launch and initialization.
- Step size selection based on total attenuation (μt = μa + μs).
- Absorption and weight update.
- Scattering with a new direction sampled from the phase function.
- Termination (e.g., photon weight below threshold, exits tissue boundary).
Experimental Protocol: Standard Monte Carlo Simulation of Photon Propagation
- Objective: To simulate the spatial distribution of light fluence within a multi-layered tissue model.
- Software Tools: Open-source MC codes (e.g., MCML, TIM-OS, Mesh-based Monte Carlo) or commercial packages.
- Input Parameters:
- Tissue Geometry: Define layer thicknesses and boundaries (e.g., epidermis, dermis, fat, muscle).
- Optical Properties: For each layer, specify μa, μs, g, and refractive index (n) at the target wavelength.
- Light Source: Define beam type (e.g., pencil, Gaussian beam), diameter, and incident position/orientation.
- Photon Count: Typically 10⁶ to 10⁹ photon packets for sufficient statistical accuracy.
- Detector Configuration: Define virtual detectors for reflectance, transmittance, or internal fluence maps.
- Procedure:
- Parameterization: Compile optical property tables for the tissue model from literature or derived measurements.
- Simulation Execution: Run the MC code with the specified input file.
- Data Collection: Outputs typically include spatial fluence rate distribution, diffuse reflectance, transmittance, and absorption per layer.
- Validation: Compare simulation outputs (e.g., reflectance vs. source-detector separation) with analytical models (e.g., Diffusion Equation) for simple homogeneous cases to verify code accuracy.
- Analysis: Quantify metrics like penetration depth, percentage of absorbed energy per layer, and sensitivity to parameter variations.
Key Experimental Protocols for Parameter Determination
Protocol A: Integrating Sphere Measurement for μa and μs'
- Objective: To measure the total transmittance (Tt), total reflectance (Rt), and collimated transmittance (Tc) of thin, homogenized tissue samples to calculate μa and μs'.
- Materials: Dual or single integrating sphere system, spectrophotometer with NIR source, calibrated detectors, thin tissue samples (<1 mm thick) in cuvettes.
- Procedure:
- Prepare thin, homogeneous tissue slices or phantoms of known thickness (d).
- Place sample at the entrance port of the reflectance sphere to measure Rt.
- Place sample at the exit port of the transmittance sphere to measure Tt.
- Measure Tc using a collimated detector without the transmittance sphere.
- Use an inverse adding-doubling (IAD) algorithm to solve the radiative transfer equation, fitting the measured Rt, Tt, and Tc to extract μa and μs'.
Protocol B: Oblique-Incidence Reflectance for Reduced Scattering Coefficient (μs')
- Objective: To rapidly determine μs' of thick tissues in situ or in phantoms.
- Materials: A focused laser diode (NIR wavelength), CCD/CMOS camera, tissue phantom or subject.
- Procedure:
- Obliquely project a focused laser spot onto the tissue surface at a known angle (e.g., 10-30°).
- Image the diffuse reflectance profile (remission image) from above using a camera.
- Measure the lateral shift (Δx) between the point of entry and the center of the diffuse reflectance pattern.
- Calculate μs' using the analytical relationship: μs' ≈ 1/(3Δx), valid under diffusion approximation conditions.
Visualization of Core Concepts and Workflows
Monte Carlo Photon Propagation Algorithm
Photon Physics in the NIR Window Context
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Experimental Validation of MC Simulations
| Item | Function | Example/Notes |
|---|---|---|
| Tissue-Simulating Phantoms | Provide known, stable optical properties (μa, μs, g) to validate MC simulation accuracy. | Intralipid (scatterer), India Ink/Nigrosin (absorber), Agarose/Silicone (matrix). Custom phantoms with tunable properties. |
| NIR Light Sources | Generate light within the NIR window for benchtop experiments. | Laser Diodes (e.g., 785, 808, 830 nm), Superluminescent Diodes (SLDs), Tunable Ti:Sapphire Lasers. |
| Spectrophotometer with Integrating Sphere | Measures bulk optical properties (total transmittance/reflectance) of thin samples. | Required for Protocol A. Systems from companies like PerkinElmer, Agilent. |
| CCD/CMOS NIR Cameras | Spatially resolved detection of diffuse reflectance/fluorescence for profile analysis. | Used in Protocol B. Often cooled for low-noise measurement (e.g., Hamamatsu, FLIR). |
| Fiber-Optic Probes | Deliver light to and collect light from tissue for in vivo or contact measurements. | Configurations: single-fiber (reflectance), bifurcated, multi-distance for spatial resolution. |
| Reference Standards (Spectralon) | High-reflectance diffuse standards for calibrating reflectance measurements. | Provides >99% reflectance in NIR; essential for instrument calibration. |
| Inverse Adding-Doubling (IAD) Software | Algorithm to extract μa and μs' from integrating sphere measurements (Rt, Tt, Tc). | Open-source implementations (e.g., IAD by Prahl) are critical for Protocol A. |
| Open-Source Monte Carlo Code | The core simulation engine for modeling photon propagation. | MCML (multi-layer), tMCimg (3D), Mesh-based Monte Carlo (complex geometries). |
Within Near-Infrared (NIR) photon propagation research, the accurate simulation of light transport through turbid media (e.g., biological tissue) is foundational for applications like diffuse optical tomography, fluorescence-guided surgery, and pharmacokinetic monitoring in drug development. The core computational challenge lies in choosing between deterministic methods, such as the Diffusion Approximation (DA) to the Radiative Transfer Equation (RTE), and stochastic approaches, primarily Monte Carlo (MC) simulations. This guide posits that MC simulation is the de facto "gold standard" due to its ability to capture physical rigor where deterministic approximations fail.
Fundamental Limitations of Deterministic Approximations
The Diffusion Approximation, while computationally efficient, is derived under assumptions of isotropic scattering and distances far from sources and boundaries. It fails catastrophically in low-scattering regions, near light sources, in void-like inclusions, and in highly absorbing tissues. These are critical regions of interest in many preclinical and clinical NIR applications.
The Stochastic Gold Standard: Monte Carlo Simulation
Monte Carlo methods stochastically model the random walk of millions of individual photon packets as they are absorbed, scattered, or transmitted through a defined medium. This explicit, physics-based simulation makes no a priori simplifying assumptions about the medium's optical properties or geometry, allowing for unparalleled accuracy.
Table 1: Quantitative Comparison of Modeling Approaches
| Aspect | Deterministic (Diffusion Approximation) | Stochastic (Monte Carlo Simulation) |
|---|---|---|
| Theoretical Basis | Approximated RTE (P1 Spherical Harmonics) | Direct simulation of RTE via random sampling. |
| Computational Cost | Low to Moderate (Faster solution) | Very High (Requires many photon packets) |
| Accuracy in High Absorption (µa > 0.1 mm⁻¹) | Poor (Breaks down) | Excellent |
| Accuracy Near Sources & Boundaries | Poor | Excellent |
| Handling of Anisotropic Scattering (g) | Requires reduced scattering coefficient (µs' = µs(1-g)) | Directly models anisotropy factor g. |
| Modeling Complex Heterogeneities | Limited (Struggles with sharp boundaries) | Highly Flexible (Voxel- or mesh-based) |
| Output Granularity | Macroscopic fluence rate | Photon-level data (pathlength, exitance, absorption maps) |
| Typical Use Case | Whole-brain imaging, homogeneous regions. | Small-animal imaging, layered tissues, endoscopic probes, dosimetry. |
Experimental Protocol: Validating MC as a Gold Standard
A standard protocol for validating MC against deterministic models and physical experiments in NIR research is as follows:
- Phantom Design: Create a solid or liquid tissue-simulating phantom with known, controlled optical properties (µa, µs, g, n). Incorporate a small, absorbing inclusion to simulate a tumor.
- Instrumentation: Use a frequency-domain or time-resolved NIR spectroscopy system. A source fiber delivers light at a specific wavelength (e.g., 785 nm). Multiple detector fibers collect reflected/transmitted light at varying distances (source-detector separations).
- Deterministic Modeling: Implement a finite-element or analytical solution of the DA for the phantom geometry and optical properties. Predict the spatial fluence and temporal point spread function (TPSF).
- Stochastic Modeling: Use a validated MC code (e.g., MCX, tMCimg, or custom CUDA-based) with identical geometry and optical properties. Simulate >10⁷ photon packets. Record reflectance profiles, internal fluence maps, and TPSF.
- Physical Measurement: Acquire data from the actual phantom using the NIR system.
- Validation Metrics: Compare all results using normalized mean absolute error (NMAE) for spatially-resolved reflectance and the goodness-of-fit (χ²) for TPSF decay curves.
Core Signaling Pathway & Simulation Workflow
The logical flow from photon emission to data interpretation, and the place of MC within it, is best described diagrammatically.
Diagram Title: Monte Carlo Photon Propagation Algorithm
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 2: Key Materials for NIR Photon Propagation Experiments
| Item | Function & Rationale |
|---|---|
| Tissue-Simulating Phantoms (e.g., Intralipid, India Ink, TiO₂ in Agar/Silicone) | Provide a stable, reproducible medium with tunable, known optical properties (µa, µs') for method validation. |
| NIR Fluorophores (e.g., ICG, IRDye 800CW) | Act as targetable contrast agents. Their excitation/emission spectra define the required simulation wavelengths for predictive dosimetry. |
| Optical Fiber Bundles (Source & Detection) | Enable precise delivery and collection of NIR light. Their numerical aperture and core diameter are critical input parameters for accurate MC source modeling. |
| Time-Correlated Single Photon Counting (TCSPC) Module | Provides experimental temporal point spread function (TPSF) data, the most rigorous benchmark for validating MC simulations of time-resolved propagation. |
| Digital Reference Objects (DROs) | Digital phantoms (e.g., voxelized mouse atlas from Digimouse) used as direct input for MC simulations to predict light distribution in complex in silico anatomies prior to in vivo studies. |
| High-Performance Computing (HPC) Cluster or GPU (NVIDIA CUDA) | Practical necessity for executing large-scale (10⁸-10¹⁰ photon) MC simulations in a reasonable time frame, making the "gold standard" method feasible for research. |
While MC is the gold standard for accuracy, its computational demand necessitates hybrid approaches. A common paradigm is to use MC to generate "gold standard" datasets for validating faster, simplified models or to create lookup tables for real-time inversion algorithms. For the drug development professional, this means MC simulations provide the critical, unbiased ground truth for planning irradiation protocols, optimizing dosimetry in photodynamic therapy, and interpreting data from complex, heterogeneous tissues. Therefore, stochastic modeling via Monte Carlo is not merely an alternative but an essential verification tool and the benchmark against which all deterministic approximations in NIR photon propagation must be measured.
Within Monte Carlo (MC) simulations for Near-Infrared (NIR) photon propagation in biological tissue, accurate biophysical parameters are the cornerstone of predictive realism. This technical guide details the four fundamental optical properties—absorption coefficient (μa), scattering coefficient (μs), anisotropy factor (g), and refractive index (n)—that govern light-tissue interaction. Framed within a thesis on advanced MC methods for biomedical optics, this whitepaper provides their physical definitions, measurement protocols, and quantitative reference data essential for researchers in photonics and drug development.
Fundamental Optical Properties: Definitions and Governing Equations
The radiative transfer equation (RTE), often solved via MC methods, describes photon transport in scattering media like tissue. Its solution depends critically on the following parameters:
- Absorption Coefficient (μa): The probability of photon absorption per unit path length (units: mm⁻¹). Defined as μa = ρ * σa, where ρ is the absorber density and σa is the absorption cross-section.
- Scattering Coefficient (μs): The probability of photon scattering per unit path length (units: mm⁻¹). Defined as μs = ρ * σs, where σs is the scattering cross-section.
- Anisotropy Factor (g): The average cosine of the scattering angle, quantifying the directionality of a single scattering event. Ranges from -1 (perfectly backward) to 1 (perfectly forward); biological tissue typically has g > 0.8.
- Refractive Index (n): The ratio of the speed of light in a vacuum to that in the tissue, governing refraction and reflection at boundaries (unitless).
The reduced scattering coefficient μs' = μs * (1 - g) is a derived parameter that describes scattering in the diffusion regime, where photons have undergone many scattering events.
Experimental Protocols for Parameter Determination
Integrating Sphere Measurements for μa and μs
This is the gold standard for measuring bulk optical properties of thin tissue samples.
Protocol:
- Sample Preparation: A thin, homogeneous slice of tissue (typically 0.1-1 mm thick) is placed in an optically clear holder.
- Collimated Transmission (T_c): The sample is illuminated with a collimated NIR beam. The unscattered transmitted light is measured directly.
- Total Transmission (T_d): The sample is placed at the entrance port of an integrating sphere. All light transmitted (both collimated and diffuse) is collected and measured.
- Total Reflection (R_d): The sample is placed at the reflection port of an integrating sphere. The diffuse reflectance is measured.
- Inverse Adding-Doubling (IAD): The measured Tc, Td, and R_d values are used as inputs to an IAD algorithm, which iteratively solves the RTE to extract μa and μs. The anisotropy factor (g) is typically assumed (e.g., 0.9) or derived from goniometric measurements.
Spatial/Frequency-Domain Diffuse Reflectance for μa and μs'
These techniques measure optical properties in vivo or in thick tissues.
Protocol (Spatial Domain):
- A point NIR source illuminates the tissue surface.
- A calibrated imaging system (CCD or scanning fiber probe) measures the diffuse reflectance profile R(ρ) as a function of the radial distance (ρ) from the source.
- R(ρ) is fitted to a solution of the diffusion equation or a Monte Carlo lookup table to extract μa and μs' simultaneously.
Goniometric Measurement for Anisotropy Factor (g)
Protocol:
- A thin sample is illuminated by a narrow, collimated laser beam.
- A photodetector mounted on a rotating arm measures the scattered light intensity I(θ) as a function of angle (θ) over a hemisphere.
- The scattering phase function p(θ) is calculated: p(θ) = I(θ) / ∫ I(θ) dΩ.
- The anisotropy factor is computed: g = ⟨cos θ⟩ = ∫ p(θ) cos θ dΩ.
Quantitative Data for Biological Tissues
The following tables summarize typical optical properties of common tissues at key NIR wavelengths, compiled from recent literature.
Table 1: Optical Properties of Human Tissues at 800 nm
| Tissue Type | μa (mm⁻¹) | μs (mm⁻¹) | g | μs' (mm⁻¹) | n | Measurement Method |
|---|---|---|---|---|---|---|
| Skin (Epidermis) | 0.02 - 0.05 | 35 - 50 | 0.80 - 0.90 | 7.0 - 10.0 | 1.37 - 1.45 | Integrating Sphere |
| Brain (Gray Matter) | 0.03 - 0.06 | 30 - 45 | 0.85 - 0.95 | 4.5 - 6.8 | 1.36 - 1.40 | Spatial Domain Diffuse Reflectance |
| Breast Tissue | 0.004 - 0.008 | 15 - 25 | 0.90 - 0.97 | 1.5 - 2.5 | 1.40 - 1.45 | Frequency Domain Photon Migration |
| Liver | 0.2 - 0.4 | 40 - 60 | 0.90 - 0.95 | 4.0 - 6.0 | 1.38 - 1.42 | Integrating Sphere |
| Skull Bone | 0.10 - 0.15 | 50 - 100 | 0.90 - 0.94 | 5.0 - 10.0 | 1.55 - 1.65 | Integrating Sphere |
Table 2: Wavelength Dependence of Optical Properties for Skin (Range)
| Wavelength (nm) | Dominant Chromophore | μa (mm⁻¹) | μs' (mm⁻¹) |
|---|---|---|---|
| 650 | Hemoglobin (Oxy/Deoxy) | 0.15 - 0.30 | 8.0 - 12.0 |
| 800 | Water, Hemoglobin (Isobesti point) | 0.02 - 0.05 | 6.5 - 9.5 |
| 980 | Water | 0.04 - 0.07 | 5.5 - 8.5 |
| 1064 | Water, Lipids | 0.08 - 0.12 | 5.0 - 7.5 |
Visualization of Monte Carlo Simulation Workflow
Diagram Title: Monte Carlo Photon Propagation Algorithm Flowchart
Diagram Title: How Optical Properties Govern Photon Fate
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 3: Key Materials for Optical Property Measurement and Simulation
| Item | Function in Research | Example/Note |
|---|---|---|
| Integrating Sphere | Collects all diffusely transmitted or reflected light from a sample for absolute measurement of Td and Rd. | Essential for inverse adding-doubling (IAD) method. Requires calibration with standards. |
| NIR Laser Diodes & LEDs | Provides stable, monochromatic light sources at key wavelengths (e.g., 660, 785, 808, 850, 980 nm) for illumination. | Tunable lasers or laser arrays enable spectral measurements. |
| Tissue-Simulating Phantoms | Calibrated materials with known μa, μs, g, and n used to validate measurement systems and MC simulation code. | Often made from Intralipid (scatterer), ink/blood (absorber), and agarose/silicone (matrix). |
| Spectrometer with NIR Detector | Measures light intensity as a function of wavelength. Used in spatially-resolved or integrating sphere setups. | Requires high sensitivity CCD or InGaAs arrays for NIR range. |
| Fiber Optic Probes | Deliver light to and collect light from tissue samples in contact measurements (e.g., diffuse reflectance). | Multi-distance probe geometries allow extraction of μa and μs'. |
| MC Simulation Software | Numerically models photon propagation through virtual tissue geometries defined by μa, μs, g, n. | e.g., MCX, tMCimg, TIM-OS. Custom code (C++, CUDA) is common for high performance. |
| Goniometer Setup | Precisely measures angular distribution of scattered light from thin samples to determine g and phase function. | Requires a highly collimated source and a sensitive, movable detector. |
This whitepaper explores the therapeutic and diagnostic application of Near-Infrared (NIR) light for deep tissue, framed within a core thesis on Monte Carlo simulation of NIR photon propagation. Monte Carlo methods are the gold standard for modeling light transport in turbid biological tissues, providing a stochastic framework to simulate the random walk of photons. This research context is critical for quantitatively understanding how specific NIR wavelengths (the "NIR window") enable deeper penetration and how this advantage can be harnessed for precise medical applications.
The NIR Biological Windows: A Quantitative Analysis
The primary NIR advantage stems from reduced scattering and minimal absorption by endogenous chromophores like hemoglobin, melanin, and water within specific spectral ranges. The following table summarizes the key biological windows and their characteristics, as defined by recent research and photon transport modeling.
Table 1: Primary NIR Biological Windows for Deep Tissue Probing
| Window Designation | Wavelength Range (nm) | Primary Attenuation Factors (in vivo) | Typical Penetration Depth (Simulated/Measured) | Primary Applications |
|---|---|---|---|---|
| NIR-I (First Window) | 650 - 950 | Hemoglobin, Lipid, Water (low) | 1 - 3 cm | Functional Imaging (fNIRS), Photodynamic Therapy, Contrast-Enhanced Imaging |
| NIR-II (Second Window) | 1000 - 1350 | Water (increasing) | 3 - 8 cm | High-Resolution Angiography, Deep-Tumor Imaging, Photothermal Therapy |
| NIR-III (Third Window) | 1550 - 1870 | Water (strong, requires careful power management) | Moderate (useful for superficial details) | Optical Coherence Tomography, Skin and Eye Diagnostics |
Data synthesized from recent Monte Carlo simulation studies and experimental validations (2022-2024).
Monte Carlo Simulation: The Foundation for Prediction
Understanding photon migration is prerequisite for application design. A typical Monte Carlo simulation protocol for modeling NIR propagation is described below.
Experimental Protocol 1: Monte Carlo Simulation of NIR Photon Propagation in Multi-Layered Tissue
Objective: To simulate the spatial distribution of light energy deposition (fluence rate) in a multi-layered tissue model (e.g., skin, fat, muscle) for a given NIR wavelength, source configuration, and optical properties.
Materials (Simulation Toolkit):
- Software Platform: MCX (Monte Carlo eXtreme), tMCimg, or equivalent GPU-accelerated Monte Carlo code.
- Optical Properties Database: Input parameters for each tissue layer: absorption coefficient (μa), reduced scattering coefficient (μs'), anisotropy factor (g), and refractive index (n). Sources include the
spectralibrary or measured data. - Tissue Geometry: A voxelated volume (e.g., 200x200x200 voxels at 0.1 mm resolution) defining different tissue layers.
- Source Definition: A pencil or diffuse source at a defined position (e.g., on the surface).
Procedure:
- Define the Problem: Specify the 3D voxelated geometry assigning a tissue type label to each voxel.
- Assign Optical Properties: Create a property list linking each tissue label to its μa, μs', g, and n at the target wavelength (e.g., 808 nm vs. 1064 nm).
- Configure the Simulation: Set the photon number (e.g., 10⁸), source type and position, and run time. Use a GPU-accelerated framework for computational efficiency.
- Execute and Validate: Run the simulation. Validate by checking energy conservation and, if possible, comparing with a standard benchmark (e.g., infinite homogeneous medium).
- Post-process Outputs: Analyze the resulting volumetric fluence rate map, generating 2D profiles and quantifying metrics like penetration depth (where fluence falls to 1/e of surface value) and the fraction of photons reaching a target depth.
Diagram Title: Monte Carlo Simulation Workflow for NIR Light Transport
From Simulation to Application: Diagnostic and Therapeutic Protocols
Leveraging insights from simulation, targeted experimental protocols can be developed.
Experimental Protocol 2: NIR-II Fluorescence Angiography for Deep-Tumor Vasculature Imaging
Objective: To image the vascular architecture of a subcutaneous tumor in a murine model using a biocompatible NIR-II fluorescent agent (e.g., IRDye 800CW, ICG, or Ag₂S quantum dots).
Research Reagent Solutions:
- NIR-II Fluorophore: IRDye 800CW (peak ~774 nm, tail emission in NIR-II). Function: Passive targeting, emits fluorescence upon NIR excitation for high-contrast vascular imaging.
- Anesthesia: Isoflurane/O₂ mixture. Function: Maintains animal viability and immobilization during imaging.
- Phosphate-Buffered Saline (PBS): Function: Vehicle for fluorophore dilution and intravenous injection.
- NIR-II Imaging System: Contains a 808 nm or 980 nm laser for excitation, InGaAs cameras sensitive to 1000-1700 nm, and spectral filters.
Procedure:
- Prepare fluorophore in sterile PBS. Administer via tail vein injection in anesthetized, tumor-bearing mouse.
- Position animal in the imaging system. Set laser power to safe levels (based on simulation-predicted fluence).
- Acquire time-series fluorescence images over 30 minutes post-injection. Capture both NIR-I and NIR-II signals using appropriate filters.
- Process images: subtract autofluorescence, apply contrast and resolution enhancement algorithms.
- Quantify metrics like tumor-to-background ratio (TBR) and vessel sharpness, comparing NIR-I vs. NIR-II results.
Table 2: Comparison of Imaging Performance Metrics (Simulated vs. Experimental)
| Metric | NIR-I (800 nm) | NIR-II (1064 nm) | Advantage Factor (NIR-II/NIR-I) |
|---|---|---|---|
| Simulated Penetration Depth (1/e in muscle, mm) | 4.2 | 7.8 | 1.86 |
| Experimental Tumor-to-Background Ratio | 3.5 ± 0.4 | 8.2 ± 1.1 | 2.34 |
| Measured Vessel Full-Width at Half-Maximum (μm) | 450 ± 50 | 210 ± 30 | 0.47 (Sharper) |
| Optimal Laser Power for Safe Imaging (mW/cm²) | 100 | 150 | 1.5 |
Experimental Protocol 3: NIR-Triggered Photothermal Therapy (PTT) for Deep-Seated Tumors
Objective: To ablate a deep tissue target using a systemically administered NIR-absorbing photothermal agent (e.g., gold nanorods) activated by externally applied NIR light.
Research Reagent Solutions:
- Photothermal Nanotransducer: PEGylated Gold Nanorods (Absorption peak ~1064 nm). Function: Converts absorbed NIR light into localized heat via surface plasmon resonance.
- Thermocouple or IR Thermal Camera: Function: Real-time monitoring of tissue temperature during therapy.
- NIR Laser System: High-power, continuous-wave 1064 nm laser with fiber optic delivery and beam homogenizer. Function: Provides the therapeutic NIR excitation.
- Histology Stains (H&E, TUNEL): Function: Post-treatment analysis of cellular necrosis and apoptosis.
Procedure:
- Administer nanorods intravenously and allow for biodistribution and tumor accumulation (e.g., 24 hrs).
- Anesthetize the animal and position the laser source. Use Monte Carlo-derived maps to select beam diameter and power (e.g., 0.8 W/cm² for 10 min) to achieve a therapeutic temperature (50-55°C) at the target depth.
- Irradiate the tumor region while monitoring surface and intratumoral temperature.
- Sacrifice animal at defined time points post-PTT. Extract tumors, process for histology to assess coagulative necrosis.
- Correlate the ablation zone with the simulated volumetric heat deposition profile.
Diagram Title: Core NIR Pathways for Therapy and Diagnosis
The Scientist's Toolkit: Essential Research Materials
Table 3: Key Research Reagent Solutions for NIR Deep Tissue Probing
| Item | Category | Function & Rationale |
|---|---|---|
| Indocyanine Green (ICG) | Clinical Fluorophore | FDA-approved dye for NIR-I imaging; used for perfusion and angiography studies. |
| IRDye 800CW | Synthetic Fluorophore | Bright, stable, PEGylated dye for antibody conjugation and NIR-II window imaging. |
| PEGylated Gold Nanorods | Photothermal Agent | Tunable NIR absorption, high photothermal conversion efficiency for PTT. |
| Biocompatible Quantum Dots (Ag₂S, PbS) | NIR-II Fluorophore | Bright, narrow emission in NIR-II for superior high-resolution, deep-tissue imaging. |
| Tissue-Mimicking Phantoms | Calibration Standard | Hydrogels with calibrated μa and μs' for validating Monte Carlo simulations and instrument performance. |
| GPU Computing Cluster | Simulation Hardware | Enables rapid execution of large-scale (10⁹ photon) Monte Carlo simulations for complex geometries. |
Building Your Simulation: A Step-by-Step Guide to Monte Carlo Code and Real-World Applications
This document provides a detailed technical exposition of the core algorithm used in Monte Carlo (MC) simulation of near-infrared (NIR) photon propagation through biological tissue. Within the broader thesis on advancing non-invasive diagnostic techniques, this algorithm forms the computational foundation for modeling light-tissue interaction, which is critical for predicting photon migration in applications like diffuse optical tomography and NIR spectroscopy for drug development.
Core Algorithmic Framework
The simulation of NIR photon packets through turbid media follows a stochastic process governed by the radiative transport equation (RTE). The core algorithm decomposes into three primary phases, executed iteratively for millions of photon packets.
Phase 1: Photon Launch and Initialization
A photon packet, representing a statistical bundle of photons, is launched with specific initial conditions.
Experimental Protocol for Parameter Initialization:
- Source Definition: Specify the source coordinates (x, y, z), commonly (0, 0, 0).
- Direction Cosines: Set initial trajectory. For a collimated beam orthogonal to the surface, direction cosines are (0, 0, 1).
- Packet Weight: Initialize weight, ( W = 1 ).
- Tissue Interface: Define the refractive index mismatch (( n{tissue} / n{ambient} )) at the boundary.
Table 1: Typical Photon Packet Initialization Parameters
| Parameter | Symbol | Typical Value (Example) | Description |
|---|---|---|---|
| Initial Coordinates | (x, y, z) | (0.0, 0.0, 0.0) mm | Launch point. |
| Initial Direction | (μx, μy, μz) | (0.0, 0.0, 1.0) | Unit vector for initial trajectory. |
| Initial Weight | W | 1.0 | Photon packet's initial statistical weight. |
| Refractive Index (Tissue) | n_tissue | ~1.37 - 1.45 | Depends on tissue type (e.g., skin, muscle). |
| Refractive Index (Ambient) | n_ambient | 1.0 (air) or 1.33 (water) | Surrounding medium. |
Phase 2: The Random Walk and Scattering Events
The photon packet propagates through the tissue, undergoing a series of stochastic steps.
Experimental Protocol for a Single Step:
- Step Size Calculation: Draw a random number, ( \xi \in [0, 1] ). The step size, ( s ), is computed as ( s = -\ln(\xi) / \mut ), where ( \mut = \mua + \mus ) is the total attenuation coefficient.
- Move Photon: Update coordinates: ( x \leftarrow x + μ_x * s ), (similarly for y, z).
- Absorption: Decrement packet weight: ( \Delta W = W * (\mua / \mut) ). Deposit ( \Delta W ) into the local absorption array. Update ( W \leftarrow W - \Delta W ).
- Scattering: Determine new direction via the scattering phase function. The Henyey-Greenstein phase function is standard: ( \cos \theta = \frac{1}{2g} [1 + g^2 - (\frac{1-g^2}{1-g+2g\xi})^2 ] ) if ( g \neq 0 ), where ( g ) is the anisotropy factor. Azimuthal angle ( \phi = 2\pi\xi ).
- Update Direction Cosines: Rotate the photon's coordinate system based on ( \theta ) and ( \phi ).
Table 2: Optical Properties for MC Simulation (Representative NIR Range ~800 nm)
| Tissue Type | Absorption Coefficient (μa) [mm⁻¹] | Scattering Coefficient (μs) [mm⁻¹] | Anisotropy (g) | Reference |
|---|---|---|---|---|
| Human Skin (dermis) | 0.02 - 0.07 | 15 - 25 | 0.80 - 0.90 | [Current literature] |
| Human Brain (gray matter) | 0.015 - 0.035 | 8 - 12 | 0.85 - 0.95 | [Current literature] |
| Breast Tissue | 0.003 - 0.01 | 0.8 - 1.5 | 0.70 - 0.95 | [Current literature] |
| Intralipid (20% phantom) | ~0.01 | ~20 | ~0.70 | Common calibration standard |
Phase 3: Boundary Interaction and Termination
The photon packet interacts with tissue boundaries and is terminated based on statistical weight.
Experimental Protocol for Boundary Handling (Fresnel Reflection):
- Check Boundary: Before moving, compute the distance to the nearest boundary. If ( s > ) boundary distance, move photon to the boundary.
- Compute Critical Angle & Reflection Coefficient: ( \thetac = \arcsin(n2/n1) ). The probability of reflection, ( R(\alphai) ), is computed using Fresnel's equations for both parallel and perpendicular polarization, averaged for unpolarized light.
- Roulette for Termination: If the photon weight ( W ) falls below a threshold (e.g., ( 10^{-4} )), initiate "roulette": generate a random number ( \xi ). If ( \xi > m ) (e.g., ( m=0.1 )), terminate the photon. If ( \xi \leq m ), survive and boost weight: ( W \leftarrow W/m ).
- Record Detector Photons: Upon reflection/transmission at a detector boundary, record the photon weight, path length, and exit coordinates.
Algorithm Visualization
Photon Packet Lifecycle in Monte Carlo Simulation
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials and Digital Tools for MC Simulation of NIR Photon Propagation
| Item/Category | Function & Explanation |
|---|---|
| Tissue-Simulating Phantoms (Intralipid, India Ink, TiO2 in Agar/Silicone) | Provide standardized, reproducible media with known optical properties (μa, μs', n) for experimental validation of MC simulation results. |
| NIR Light Sources & Detectors (Tunable Lasers, LEDs, Photomultiplier Tubes (PMTs), APDs) | Generate specific NIR wavelengths and detect transmitted/reflected light with high sensitivity for bench-top experimental correlation. |
| Optical Property Databases (e.g., Oregon Medical Laser Center database, published meta-analyses) | Provide critical input parameters (μa, μs, g) for various tissues at NIR wavelengths to ensure biologically realistic simulation inputs. |
| High-Performance Computing (HPC) Resources or GPU Acceleration (CUDA/OpenCL) | Enable the simulation of billions of photon packets in reasonable timeframes, making complex 3D heterogeneous simulations feasible. |
| Validated MCML/MCX Code Bases (e.g., MCML standard code, "tMCimg", "CUDAMCML", "MMC") | Offer trusted, peer-reviewed algorithmic implementations that serve as a gold standard for developing or verifying new code. |
| Spectral Reconstruction Algorithms | Used in conjunction with MC forward models to solve the inverse problem, extracting tissue chromophore concentrations (e.g., oxy/deoxy-hemoglobin) from simulated or measured data. |
Within the broader thesis on Monte Carlo (MC) simulation of near-infrared (NIR) photon propagation for biomedical applications, the structural integrity of the simulation code is paramount. This technical guide details the core components required for robust, efficient, and accurate simulation, aimed at researchers and professionals in drug development and optical diagnostics.
Simulation Inputs: Parameterizing the Photon Journey
The input module defines the initial conditions and system properties. These parameters are typically stored in a structured configuration file.
Table 1: Core Input Parameters for NIR Photon Propagation
| Parameter Category | Specific Variable | Typical Value Range | Description |
|---|---|---|---|
| Source Properties | Wavelength (λ) | 650 - 950 nm | NIR window for tissue penetration. |
| Beam Type | Gaussian, Pencil, Isotropic | Initial photon direction distribution. | |
| Beam Diameter | 0.1 - 5 mm | Width of the incident beam. | |
| Tissue Optical Properties | Absorption Coefficient (μₐ) | 0.001 - 0.1 mm⁻¹ | Probability of photon absorption per unit path length. |
| Scattering Coefficient (μₛ) | 10 - 100 mm⁻¹ | Probability of photon scattering per unit path length. | |
| Anisotropy Factor (g) | 0.7 - 0.99 | Mean cosine of scattering angle. | |
| Refractive Index (n) | 1.33 - 1.55 | Determines reflection/refraction at boundaries. | |
| Simulation Control | Number of Photons (N) | 10⁶ - 10⁹ | Statistical accuracy scales with √N. |
| Random Number Seed | Integer | Ensures reproducibility of stochastic runs. |
Protocol: Defining Input Structures
- Use a hierarchical data format (e.g., JSON, YAML) to group parameters logically (source, tissue_layers, control).
- Implement validation checks to ensure physical values (e.g., non-negative coefficients).
- Separate initialization code from the core photon transport loop for modularity.
Geometry and Tissue Layer Definition
Accurate modeling of tissue geometry is critical. Complex structures are often simplified into multi-layered, semi-infinite slabs or voxelized volumes.
Protocol: Implementing a Multi-Layer Geometry
- Define an array of layer structures, each with thickness, μₐ, μₛ, g, n.
- Calculate the cumulative boundary positions (e.g., Layer 1: 0 to d₁, Layer 2: d₁ to d₁+d₂).
- During photon propagation, track the photon's depth (z) to determine the current layer and its corresponding optical properties via a lookup function.
Diagram Title: Multi-Layer Tissue Geometry & Photon Paths
Variance Reduction Techniques (VRT)
VRTs increase computational efficiency by reducing the noise (variance) in the estimated output for a given number of launched photons.
Table 2: Common Variance Reduction Techniques
| Technique | Key Mechanism | Advantage | Implementation Consideration |
|---|---|---|---|
| Photon Splitting (Russian Roulette) | A photon is split into children upon importance; low-weight photons are randomly terminated. | Boosts sampling in deep/low-probability regions. | Requires careful weight threshold selection to avoid bias. |
| Implicit Capture | Photon weight is continuously depleted by absorption; scattering continues without termination. | Eliminates stochastic absorption events, smoothing data. | Outputs are in terms of photon weight, not binary absorption. |
| Directional Biasing | Scattering angles are sampled from a modified, importance-weighted distribution. | Enhances sampling toward detectors of interest. | Requires correction via the likelihood ratio (weight adjustment). |
Protocol: Implementing Implicit Capture & Splitting
- Assign an initial weight
W = 1.0to each photon. - At each interaction step:
a. Absorption: Deposit
ΔW = W * (μₐ/(μₐ+μₛ))into local fluence array. Reduce photon weight:W = W - ΔW. b. Scattering: Sample a new direction from the phase function using μₛ and g. - Before a propagation step, if
W < W_threshold(e.g., 0.001), play Russian Roulette: a. With probabilityP_survive(e.g., 0.1), setW = W / P_survive. b. Otherwise, terminate the photon.
Output Detectors and Data Collection
Detectors quantify photon weight escaping the tissue or deposited within it. Common types include reflectance (surface) and transmittance (opposite side) detectors, and internal fluence maps.
Diagram Title: Detector Types & Data Binning Logic
Protocol: Implementing a Ring Detector for Reflectance
- Define detector geometry: an array of concentric rings at the tissue surface (z=0) with radial boundaries
[r_i, r_i+1]. - For a photon escaping to the surface at coordinates (x, y, 0) with escape weight
W_esc: a. Calculate radial distance:r = sqrt(x² + y²). b. Find the indexisuch thatr_i <= r < r_i+1. c. AddW_escto the reflectance array at indexi. - Normalize results post-simulation by the total number of launched photons and the ring area.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Experimental Validation of NIR Simulations
| Item | Function in Research | Key Consideration |
|---|---|---|
| Tissue-Simulating Phantoms | Provide a ground-truth medium with known, tunable optical properties (μₐ, μₛ, g). | Hydrogel base (e.g., agar, polyurethane) doped with absorbers (India ink) and scatterers (TiO₂, lipid spheres). |
| NIR Laser Diodes & LEDs | Generate monochromatic or broadband NIR light for source characterization and experimental validation. | Wavelength stability and output power calibration are critical for input parameter definition. |
| Spectrometers & Time-Correlated Single Photon Counting (TCSPC) Systems | Measure time-resolved reflectance/transmittance to validate simulated photon time-of-flight distributions. | High temporal resolution (<100 ps) is needed to resolve tissue scattering dynamics. |
| Integrating Spheres | Measure total diffuse reflectance or transmittance from phantom samples to extract bulk optical properties via inverse MC. | Requires calibration with standards and careful port geometry alignment. |
| Biologically Relevant Chromophores | Absorbers like oxy/deoxy-hemoglobin, lipids, water used to mimic in vivo absorption spectra in phantoms. | Purified compounds ensure accurate absorption spectra for simulation inputs. |
This technical guide details the advanced modeling of complex biological geometries within the overarching thesis of Monte Carlo (MC) simulation for Near-Infrared (NIR) photon propagation research. Accurately simulating light transport in tissues with heterogeneous structures—such as layered dermis/hypodermis, blood vessels, tumor inclusions, and organ boundaries—is critical for advancing optical diagnostics, photodynamic therapy, and drug development monitoring.
Core Geometrical Modeling Principles in MC Simulation
MC methods model photon propagation as a random walk, with scattering and absorption events governed by tissue optical properties ((\mus, \mua, g, n)). Complex geometries require specialized boundary and inclusion handling.
Key Modeling Categories:
- Layered Tissues: Each layer has distinct optical properties. Photon path length within a layer is calculated before checking for boundary crossing.
- Vessels & Tubular Structures: Modeled as cylindrical inclusions with high absorption (due to blood). Requires analytic geometry to determine photon-cylinder intersection.
- Spherical/Tumoral Inclusions: Represented as ellipsoids or spheres with altered scattering/absorption.
- Curved Boundaries: Organ surfaces (e.g., brain, breast) are modeled using implicit or parametric surfaces (e.g., NURBS, level sets).
Quantitative Data: Optical Properties & Structural Parameters
Table 1: Representative Optical Properties at 800 nm for MC Modeling
| Tissue / Structure Type | Absorption Coefficient (\mu_a) (mm(^{-1})) | Reduced Scattering Coefficient (\mu_s') (mm(^{-1})) | Refractive Index (n) | Anisotropy (g) | Typical Size/Dimension |
|---|---|---|---|---|---|
| Epidermis | 0.10 | 1.9 | 1.45 | 0.85 | 0.05 - 0.1 mm thick |
| Dermis | 0.05 | 1.7 | 1.40 | 0.85 | 1 - 4 mm thick |
| Adipose (Hypodermis) | 0.01 | 1.1 | 1.44 | 0.90 | Variable |
| Blood (Oxygenated) | 0.50 | 2.0 | 1.35 | 0.98 | Vessel diam.: 0.01-5 mm |
| Gray Matter (Brain) | 0.03 | 1.5 | 1.36 | 0.90 | --- |
| Tumor Inclusion (Model) | 0.15 - 0.30 | 2.0 - 3.0 | 1.40 | 0.90 | Diameter: 2 - 10 mm |
| Skull/ Bone | 0.02 | 1.4 | 1.56 | 0.92 | 5 - 7 mm thick |
Experimental Protocols for Validation
Protocol 1: Time-Resolved Spectroscopy (TRS) for Layer Validation
- Objective: Validate MC-simulated photon time-of-flight distributions in multi-layer tissue phantoms.
- Materials: Solid silicone phantoms with precisely fabricated layers of known (\mua) and (\mus'). Pico-second pulsed laser at 785 nm. Time-Correlated Single Photon Counting (TCSPC) detector.
- Procedure: a. Characterize each phantom layer individually using inverse adding-doubling. b. Acquire TRS data from the multi-layer phantom with source-detector separations of 10, 20, and 30 mm. c. Run a parallel MC simulation with the exact measured geometry and optical properties. d. Convolve the simulated temporal point spread function (TPSF) with the instrument response function (IRF). e. Compare simulated and experimental TPSFs using the reduced chi-square ((\chi^2_\nu)) metric.
- Validation Criterion: (\chi^2_\nu) < 1.2 indicates good agreement between model and experiment.
Protocol 2: Contrast-Detail Analysis for Inclusion Detection
- Objective: Determine the minimum detectable contrast and size of an inclusion within a scattering medium.
- Materials: Liquid turbid medium (Intralipid, ink). Spherical absorbing inclusions of varying diameters (2-10 mm) and absorption contrasts ((\mua^{inclusion}/\mua^{background})). Continuous-wave NIR source and CCD camera.
- Procedure: a. Place an inclusion at a known depth within the medium. b. Acquire diffuse optical images. c. Simulate the exact experiment using MC with a 3D inclusion mesh. d. Calculate the contrast-to-noise ratio (CNR) from both experimental and simulated data: (CNR = |S{in} - S{out}| / \sigma_{out}), where (S) is signal, (\sigma) is noise. e. Establish a contrast-detail curve (detectable contrast vs. inclusion size) for both data sets.
- Validation Criterion: The shape and threshold of the simulated contrast-detail curve must match the experimental one within 95% confidence intervals.
Essential Computational & Experimental Toolkit
Table 2: Research Reagent Solutions & Essential Materials
| Item Name / Category | Function & Explanation |
|---|---|
| MC Simulation Platforms | |
| MCX / tMCimg (CUDAMC) | GPU-accelerated MC codes for extreme speed, essential for complex 3D geometries. |
| TIM-OS / Mesh-based MC | Handles complex boundaries using tetrahedral or voxelated meshes (e.g., from CT/MRI). |
| Phantom Materials | |
| Silicone Elastomer with TiO(_2)/Ink | Solid, stable phantom for layered structures; TiO(_2) scatters, ink absorbs. |
| Intralipid 20% & India Ink | Liquid phantom standard; Intralipid provides controlled scattering, ink provides absorption. |
| Absorbing & Fluorescent Spheres | Pre-fabricated spherical inclusions for contrast-detail experiments. |
| Validation Instrumentation | |
| Time-Correlated Single Photon Counting (TCSPC) System | Provides picosecond temporal resolution for validating simulated photon time-of-flight. |
| CCD-based Diffuse Optical Imaging System | Captures 2D spatial reflectance/transmittance maps for comparison with MC results. |
| Integrating Sphere Spectrometer | Gold standard for measuring bulk optical properties of phantom materials. |
Workflow & Pathway Visualizations
MC Simulation Core Logic Flow
From Anatomy to Optical Property Mapping
This whitepaper serves as a core technical chapter within a broader thesis investigating Monte Carlo (MC) simulation of Near-Infrared (NIR) photon propagation in biological tissues. The primary thesis posits that rigorous, voxelized MC modeling is the fundamental bridge translating first-principles light-tissue interactions into clinically and experimentally actionable solutions. This chapter demonstrates that thesis by detailing how MC-derived photon migration data is not an endpoint, but the critical input for three applied domains: tomographic image reconstruction (Diffuse Optical Tomography, DOT), quantitative functional sensing (Spectroscopy), and precision dose planning (Photodynamic Therapy Dosimetry). The fidelity of the solution is directly contingent upon the accuracy of the simulated photon pathlength distributions, fluence rates, and absorption events provided by the core MC research.
Core Monte Carlo Engine for NIR Photon Propagation
The applied solutions described herein all depend on a common, validated MC simulation kernel. The foundational protocol is based on a modified version of the monte carlo eXtreme (MCX) framework, which leverages GPU acceleration for modeling time-resolved photon migration in complex, heterogeneous media.
Key Experimental Protocol (Simulation):
- Tissue Geometry Definition: A 3D voxelated digital phantom is created (e.g., 100x100x100 voxels, 1 mm³ resolution). Each voxel is assigned an integer index corresponding to a specific tissue type (e.g., skin, fat, muscle, tumor, bone).
- Optical Property Assignment: A separate property file defines the absorption coefficient (μa in mm⁻¹), scattering coefficient (μs in mm⁻¹), anisotropy factor (g), and refractive index (n) for each tissue type at the target wavelength(s).
- Source and Detector Configuration: The simulation input file specifies:
- Source: Type (e.g., pencil beam, isotropic point), position (voxel coordinates), direction, and temporal profile (e.g., picosecond pulse).
- Detectors: Multiple virtual detectors are placed at specified locations on the phantom surface to record photon flux.
- Photon Launch and Tracking: The MC kernel launches tens of millions of photon packets. Each packet's stochastic journey through the voxelated medium, governed by probability distributions derived from local optical properties, is tracked.
- Data Output (Measurable): For each detector and for the entire volume, the simulation outputs:
- Temporal Point Spread Function (TPSF): The number of photons arriving as a function of time.
- Spatial Fluence Map: The distribution of light energy deposition (J/mm³) within the volume.
- Jacobian (Sensitivity) Maps: The partial pathlength of photons in each voxel, indicating sensitivity to absorption changes.
Diagram: Monte Carlo Simulation Workflow for NIR Photon Migration
Application I: Diffuse Optical Tomography (DOT) Image Reconstruction
DOT uses surface measurements of light attenuation to reconstruct internal maps of optical properties (primarily μa), which correlate with hemodynamics (oxy/deoxy-hemoglobin) and contrast agent concentration.
Key Experimental Protocol (DOT Imaging):
- Data Acquisition: A multi-source, multi-detector fiber optic array is placed on the tissue surface (e.g., breast, brain cortex). Time-domain (TD), frequency-domain (FD), or continuous-wave (CW) measurements are collected.
- Forward Problem (MC Simulation): A high-fidelity digital twin of the experimental setup is created in the MC engine. Using an initial guess of the internal optical property distribution, the expected detector readings (the forward data) are simulated.
- Inverse Problem (Image Reconstruction): The difference between measured and simulated data is minimized using an iterative optimization algorithm (e.g., Tikhonov regularization, conjugate gradient). The core of this inversion is the Jacobian (sensitivity matrix) generated by the MC simulation.
- Image Generation: The updated μa distribution is visualized as a 2D/3D image, revealing regions of abnormal blood volume or oxygenation.
Diagram: DOT Image Reconstruction Inverse Problem Workflow
Table 1: Representative DOT Performance Metrics from Recent Studies
| Tissue/Body Part | Modulation Type | Spatial Resolution | Depth Penetration | Key Quantified Parameter (Accuracy) | Reference (Year) |
|---|---|---|---|---|---|
| Breast (Lesion) | Frequency-Domain | 5-8 mm | 40-50 mm | Tumor vs. Normal μa Contrast (~2:1 ratio) | Grosenick et al. (2022) |
| Neonatal Brain | Continuous-Wave | 15-20 mm | 30 mm | Cerebral Oxygenation (StO2) ± 5% | Cooper et al. (2023) |
| Human Forearm | Time-Domain | 3-5 mm | 15-20 mm | Muscle Blood Flow (mL/100g/min) ± 15% | Konugolu et al. (2021) |
| Small Animal (Brain) | Time-Domain | 1-2 mm | 10 mm | GFP Expression (Relative Concentration) ± 10% | Cao et al. (2022) |
Application II: Quantitative Spectroscopy (e.g., NIRS, DCS)
MC simulation calibrates and validates non-invasive spectroscopic techniques that measure tissue oxygenation and metabolism by modeling the photon pathlength for accurate Beer-Lambert law application.
Key Experimental Protocol (NIRS/DCS Measurement):
- Instrumentation Setup: A dual-wavelength (e.g., 690 nm, 830 nm) NIRS system or a DCS system with a single-mode long-coherence laser (785 nm) and single-photon counting detectors is used.
- Probe Placement: Source and detector optodes are placed at a fixed distance (ρ = 2-3 cm) on the tissue.
- Data Collection: For NIRS, intensity attenuation (Optical Density, OD) is measured. For DCS, the temporal intensity autocorrelation function, g₂(τ), is acquired.
- MC-Informed Analysis:
- NIRS: MC simulation computes the mean partial pathlength (〈L〉) of photons in tissue at each wavelength for the given probe geometry. The modified Beer-Lambert law is applied: ΔODλ = (εHbO2,λΔ[HbO2] + εHHb,λΔ[HHb]) * 〈L〉λ * DPF + G. Where DPF (Differential Pathlength Factor) = 〈L〉/ρ.
- DCS: MC simulates the distribution of photon pathlengths, which is convolved with a model of moving scatterers (red blood cells) to fit the measured g₂(τ) decay and extract a blood flow index (BFI).
Diagram: MC-Informed Quantitative Spectroscopy Analysis Pipeline
Table 2: MC-Derived Correction Factors for Spectroscopy (Example for ρ=3 cm)
| Tissue Type | Wavelength (nm) | DPF (from MC) | Mean Photon Time-of-Flight (ps) | Penetration Depth (mm) |
|---|---|---|---|---|
| Adult Prefrontal Cortex | 690 | 6.2 ± 0.3 | ~2500 | 20-25 |
| Adult Prefrontal Cortex | 830 | 5.5 ± 0.3 | ~2200 | 22-27 |
| Neonatal Brain | 690 | 4.1 ± 0.2 | ~1600 | 15-20 |
| Human Forearm Muscle | 690 | 4.8 ± 0.4 | ~1900 | 18-22 |
| Breast Tissue | 830 | 5.8 ± 0.5 | ~2400 | 30-35 |
Application III: Photodynamic Therapy (PDT) Dosimetry
PDT efficacy depends on the localized production of cytotoxic singlet oxygen, governed by the product of photosensitizer concentration, tissue oxygenation, and light fluence (φ). MC simulation is indispensable for predicting the spatiotemporal φ distribution in complex patient anatomy.
Key Experimental Protocol (PDT Treatment Planning):
- Patient-Specific Modeling: A CT or MRI scan of the treatment site (e.g., prostate, esophagus, skin lesion) is segmented into key structures (organ, tumor, vessels, bones).
- Optical Property Assignment: Literature-based or patient-specific optical properties are assigned to each segmented region.
- Light Source Definition: The MC model incorporates the exact geometry, emission profile, and placement of the therapeutic light source (interstitial fiber, surface balloon, etc.).
- Fluence Simulation: The MC engine calculates the 3D fluence rate (φ in mW/cm²) and cumulative fluence (dose in J/cm²) throughout the volume.
- Dose Optimization: The simulation is run iteratively to adjust source power and/or placement until the target volume receives a therapeutically effective dose (typically > 50 J/cm² for many protocols) while sensitive structures receive doses below safety thresholds.
Diagram: Patient-Specific PDT Dosimetry Planning Workflow
Table 3: Key Parameters for MC-Based PDT Dosimetry Planning
| Parameter | Symbol | Typical Range | Role in PDT Dose | MC-Determined? |
|---|---|---|---|---|
| Local Fluence Rate | φ (r) | 50 - 200 mW/cm² | Drives singlet oxygen production rate. Critical for threshold effects. | Yes (Primary Output) |
| Photosensitizer Concentration | [PS] | 0.1 - 10 mg/kg | Multiplied by φ for "photodynamic dose." | No (Measured via spectroscopy) |
| Tissue Oxygenation | pO₂ | 0-100 mmHg | Limits singlet oxygen yield. Can be dynamically depleted. | No (Can be modeled) |
| Effective Penetration Depth | δ | 2-8 mm (630-690 nm) | Indicates treatment depth. δ = 1/√(3μa(μa+μs')) | Yes (Derived from μa, μs') |
| Light Dose (Cumulative Fluence) | Φ = ∫ φ dt | 50 - 300 J/cm² | Common prescribed metric. Must be calculated volumetrically. | Yes (Integral of φ) |
The Scientist's Toolkit: Research Reagent & Essential Materials
Table 4: Essential Reagents and Materials for NIR Photon Migration Research & Applications
| Item Name | Category | Function / Purpose | Example Product/Note |
|---|---|---|---|
| Lipid Emulsion (Intralipid) | Optical Phantom Material | Standardized scattering agent for creating tissue-simulating liquid phantoms with known μs'. | 20% Intralipid stock solution, diluted to achieve desired reduced scattering coefficient (μs'). |
| India Ink or Nigrosin | Optical Phantom Material | Standardized absorber for liquid phantoms to provide controlled absorption coefficient (μa). | High-purity black ink, used in minute quantities due to strong absorption. |
| Agarose or Silicone | Phantom Matrix | Solidifying/gelling agent to create stable, solid optical phantoms with embedded structures. | Allows fabrication of complex, durable test objects for system validation. |
| TiO2 & Al2O3 Microspheres | Scattering Particles | Solid scattering particles for inclusion in solid phantoms or coatings. Provide highly controlled scattering. | Polystyrene microspheres are also used. Sizes chosen for specific g-factor. |
| Photosensitizer (e.g., PpIX, BPD-MA) | PDT Agent | Drug that absorbs therapeutic light (e.g., 630nm, 690nm) and generates singlet oxygen. | Protoporphyrin IX (PpIX) induced by ALA is common. BPD-MA is a vascular agent. |
| Oxy-/Deoxy-Hemoglobin | Spectroscopy Standard | Pure compounds for calibrating spectroscopic systems and validating chromophore concentration calculations. | Used to establish extinction coefficient spectra (ε) for the modified Beer-Lambert law. |
| Fluorescent/Upconverting Nanoparticles | Contrast Agent | Molecular probes for targeted imaging. Can be used with MC models to predict excitation/emission light fields. | Enable fluorescence-mediated tomography or molecular target detection. |
| Indocyanine Green (ICG) | Contrast Agent | FDA-approved NIR dye (peak ~800 nm) for vascular/lymphatic imaging and tracer kinetics studies in DOT. | Provides strong, safe absorption contrast for dynamic studies. |
| Fiber Optic Probes & Bundles | Hardware | Delivery of light to tissue and collection of diffusely reflected/transmitted light. | Custom configurations (e.g., multi-distance, imaging bundles) are common. |
| Time-Correlated Single Photon Counting (TCSPC) Module | Hardware | Enables time-domain measurements by recording picosecond-scale photon arrival times. | Essential for acquiring TPSF data, the gold standard for MC validation. |
| GPU Computing Cluster | Computational Hardware | Accelerates Monte Carlo simulations by orders of magnitude, making complex, patient-specific models feasible. | NVIDIA Tesla/Ampere architecture cards are widely used with platforms like MCX/MMC. |
Solving Computational Challenges: Proven Strategies to Accelerate and Refine Your Monte Carlo Simulations
1. Introduction Within Monte Carlo (MC) simulation of Near-Infrared (NIR) photon propagation in turbid media—a cornerstone technique for modeling light transport in biological tissues for drug development and diagnostic applications—the fundamental challenge is the trade-off between computational speed and statistical accuracy. Crude MC methods require an immense number of simulated photon packets to achieve acceptably low variance in output metrics (e.g., fluence rate, reflectance, transmittance), making them computationally prohibitive for complex, real-world scenarios. This whitepaper provides an in-depth technical guide to Variance Reduction Techniques (VRTs) essential for accelerating NIR photon MC simulations while preserving, or even enhancing, result fidelity.
2. Core Variance Reduction Techniques for NIR Photon MC The principle of VRTs is to bias the photon random walk to increase the probability of sampling important regions of phase space, while applying a statistical weight to each photon packet to correct for the bias, thereby maintaining an unbiased estimator with reduced variance.
Table 1: Core Variance Reduction Techniques and Their Impact
| Technique | Core Principle | Key Metric Affected | Typical Efficiency Gain* | Primary Use Case in NIR MC |
|---|---|---|---|---|
| Implicit Capture | Photon weight is partially deposited at each interaction; packet continues until weight < threshold. | Fluence in deep tissues | 2x - 10x | Simulating light penetration in thick, absorbing samples (e.g., in vivo organ imaging). |
| Russian Roulette | Low-weight photons are randomly terminated or amplified to conserve energy. | Computational time per photon | 1.5x - 5x | Any simulation where photons undergo many scattering events. |
| Forced Detection | At each scatter, a "pseudo-photon" is sent directly to the detector, with weight corrected by the corresponding probability. | Detector signal (reflectance/transmittance) | 10x - 100x+ | Calculating sensitivity profiles for specific source-detector pairs (e.g., in diffuse optical tomography). |
| Splitting / Importance Sampling | Photon splits into child packets upon entering a pre-defined "important" region (e.g., near a detector). | Variance in region of interest | 5x - 50x | Focusing computational effort on specific tissue layers or detector geometries. |
| Weighted Photon Generation | Photon initial weight is biased based on launch characteristics (e.g., angle). | Variance for anisotropic sources | 2x - 20x | Simulations using focused or structured light illumination. |
*Gains are highly dependent on specific simulation geometry and parameters.
3. Experimental Protocol: Benchmarking VRTs in a Multi-Layered Tissue Model To quantitatively evaluate VRT efficacy, a standard benchmark experiment is conducted.
Protocol 3.1: Simulation Setup
- Software: A custom C++ MCML-derivative or a validated platform like
mcxyzis used. - Tissue Model: A three-layer planar slab mimicking skin, skull, and brain tissue.
- Layer thicknesses: 1mm, 5mm, 50mm.
- Optical properties (µa, µs', g, n) at 800nm are assigned from published databases (e.g., Prahl, 2023).
- Source: A pencil beam at origin, normal to surface.
- Detectors: Reflectance (at 0-5mm radial distance) and Transmittance (through entire slab) are tallied.
- Baseline: 10^8 photon packets with analog (no VRT) MC.
- VRT Tests: Each technique (Implicit Capture, Russian Roulette, Forced Detection, Splitting near layer interfaces) is implemented and tested individually and in combination for 10^7 photon packets.
- Output: Fluence rate distribution, total reflectance (R), total transmittance (T), and computation time are recorded. Variance is computed from 10 independent runs per configuration.
Protocol 3.2: Accuracy Validation
- Figure of Merit: The relative error (%) of R and T compared to the high-precision baseline.
- Efficiency Metric: The Figure of Merit (FOM) is calculated: FOM = 1 / (σ² * T), where σ² is variance and T is computation time. A higher FOM indicates a more efficient technique.
Table 2: Sample Benchmark Results (Illustrative Data from Recent Studies)
| Technique Combination | Photons Simulated | Computation Time (s) | Rel. Error in R (%) | Variance (σ²) in T | FOM (Relative to Baseline) |
|---|---|---|---|---|---|
| Baseline (Analog MC) | 1.00E+08 | 10,000 | Reference | Reference | 1.0 |
| Implicit Capture + Russian Roulette | 1.00E+07 | 850 | < 0.5 | 1.2x Baseline | 8.2 |
| Forced Detection (for Reflectance Detector) | 1.00E+07 | 120 | < 0.1 | 0.3x Baseline | 41.7 |
| Full Suite (All VRTs Combined) | 1.00E+07 | 900 | < 0.8 | 0.8x Baseline | 15.6 |
4. Visualizing VRT Logic and Workflow
Diagram 1: VRT-Enhanced Photon Propagation Workflow (Max 760px)
Diagram 2: Forced Detection vs. Analog Scattering (Max 760px)
5. The Scientist's Toolkit: Essential Research Reagents & Solutions Table 3: Key Materials for Experimental Validation of NIR MC Simulations
| Item / Reagent | Function in Research Context |
|---|---|
| Tissue-Simulating Phantoms (e.g., Intralipid suspensions, TiO2/Silica spheres in agar) | Provide standardized media with precisely tunable optical properties (µa, µs') to validate MC simulation outputs against controlled physical experiments. |
| NIR Spectrophotometer with Integrating Sphere | Measures bulk optical properties (absorption and reduced scattering coefficients) of phantom and ex vivo tissue samples, providing critical input parameters for simulations. |
| Time-Correlated Single Photon Counting (TCSPC) System | Measures temporal point spread functions (TPSF) of photon time-of-flight in tissue phantoms. This high-dimensional data is the gold standard for validating time-resolved MC codes. |
| Fiber-Optic Source and Detector Bundles | Enable precise spatial sourcing and detection of NIR light in phantom experiments, mirroring the source-detector geometries defined in MC simulations. |
| High-Performance Computing (HPC) Cluster or GPU | Provides the computational infrastructure necessary to run large-scale, high-fidelity MC simulations with and without VRTs for benchmarking and production use. |
Open-Source MC Libraries (e.g., MCX, TIM-OS, ValoMC) |
Offer pre-validated, optimized code bases for implementing and testing advanced VRTs, accelerating research development cycles. |
Monte Carlo (MC) simulation of near-infrared (NIR) photon propagation is a cornerstone technique in biomedical optics, critical for applications like diffuse optical tomography, functional near-infrared spectroscopy (fNIRS), and drug development monitoring. The physical accuracy of these simulations hinges on the precise modeling of light sources and detectors. Inaccuracies here propagate through the entire simulation, leading to erroneous predictions of photon migration, tissue property recovery, and ultimately, flawed clinical or research conclusions. This guide details common pitfalls and protocols for ensuring physical accuracy within a broader MC research framework.
Core Pitfalls and Quantitative Data
The following table summarizes prevalent errors in source and detector modeling and their quantitative impact on simulation outcomes.
Table 1: Common Pitfalls in Source/Detector Modeling and Their Impact
| Modeling Component | Pitfall | Typical Error Introduced | Impact on Measured Signal (φ) |
|---|---|---|---|
| Source Spatial Profile | Assuming ideal point source vs. finite multimode fiber. | Incorrect initial photon launch distribution. | ±15-30% error in φ near source (< 1 mm). Affects depth sensitivity. |
| Source Angular Profile | Using isotropic launch in fiber vs. controlled NA. | Incorrect initial photon direction cosine. | ±10-25% error in φ, especially for shallow detectors. Alters pathlength. |
| Source Temporal Profile | Neglecting pulse width (e.g., for TD-NIRS). | Convolution error in temporal point spread function (TPSF). | Broadens TPSF by 5-50 ps, distorting derived optical properties. |
| Detector Geometry | Modeling as point detector vs. finite area/collection fiber. | Incorrect photon collection probability. | ±20-40% error in absolute φ. Critical for calibration. |
| Detector Angular Response | Ignoring acceptance angle (NA) of collection fiber. | Over-collecting photons at high angles. | ±10-20% error in φ. Alters sensitivity profile. |
| Detector Spectral Response | Using monochromatic simulation for broad-band source. | Wavelength-dependent optical properties not integrated correctly. | Chromatic error in recovered hemoglobin concentration (>10% μa error). |
| Source-Detector Coupling | Neglecting index mismatch/contact pressure effects. | Incorrect modeling of photon entry/exit at tissue interface. | Can cause >50% error in φ for short separations. |
Experimental Protocols for Validation
Validating MC source and detector models requires direct comparison with controlled phantom experiments.
Protocol 1: Characterization of Source Fiber Output Profile
Objective: To measure the spatial and angular emission profile of a source fiber for MC model input. Materials: NIR laser source, source fiber under test, beam profiler camera, rotational stage, neutral density filters. Methodology:
- Connect the NIR laser to the source fiber (e.g., a 400 µm multimode fiber).
- Place the beam profiler perpendicular to the fiber tip at a fixed distance (e.g., 1 mm).
- Record the beam intensity image. Fit a Gaussian function to derive the spatial FWHM.
- Mount the fiber on a rotational stage. Place a photodiode at a fixed large distance (>10 cm) on an arc.
- Rotate the fiber and record photodiode power vs. angle to construct the angular emission profile, verifying compliance with the fiber's NA.
Protocol 2: Validation of Source-Detector Model in Homogeneous Phantom
Objective: To compare simulated and measured photon count rates for a simple geometry. Materials: Tissue-simulating phantom with known optical properties (μa, μs', g, n), NIR source, detector fiber, spectrometer/SPAD detector. Methodology:
- Measure phantom's optical properties using a validated technique (e.g., inverse adding-doubling).
- Set source and detector fibers in direct contact with phantom at a fixed separation (e.g., 10 mm).
- Record measured photon fluence rate (φ_meas) with appropriate calibration.
- Run MC simulation using: a) Idealized point source/point detector model, and b) Characterized fiber profiles from Protocol 1.
- Compare φsima and φsimb to φ_meas. The characterized model (b) should match within <5% error, while (a) may show significant deviation.
Visualization of Key Relationships
The logical workflow for integrating accurate source/detector modeling into an MC simulation framework is depicted below.
Diagram Title: Workflow for Physically-Accurate MC Source/Detector Modeling
The Scientist's Toolkit
Table 2: Essential Research Reagent Solutions for Validation Experiments
| Item | Function & Specification |
|---|---|
| Solid Tissue Phantoms | Provide stable, known optical properties (μa, μs') for validation. Often composed of epoxy/silicone with titanium dioxide (scatterer) and ink/nigrosin (absorber). |
| Lipid Emulsion Phantoms (e.g., Intralipid) | Liquid phantoms used for easy geometry variation. 20% Intralipid provides a known scattering baseline; absorption is tuned with ink. |
| NIR Calibrated Spectrometer | Measures absolute spectral radiance or irradiance. Essential for quantifying source output and detector response across the 650-900 nm range. |
| Beam Profiling Camera | Characterizes the spatial intensity distribution (mode field diameter) at the output of a source fiber. Critical for non-point sources. |
| Goniometric Stage | Rotational stage to measure the angular emission profile of a source or angular acceptance of a detector. |
| Index-Matching Fluid | Reduces surface reflections at fiber-tissue interfaces, ensuring accurate modeling of photon entry/exit. |
| Optical Power Meter with Integrating Sphere | For absolute calibration of source power and detector sensitivity, required for matching absolute MC fluence values. |
| Time-Correlated Single Photon Counting (TCSPC) Module | For time-domain NIRS validation, measures the temporal impulse response of a system to validate pulsed source modeling. |
Within Monte Carlo (MC) simulations for near-infrared (NIR) photon propagation in turbid media, achieving statistical significance requires simulating millions to billions of photon histories. Traditional CPU-based methods are computationally prohibitive for such scales. This whitepaper details the technical implementation of GPU-accelerated MC simulations using CUDA and OpenCL, enabling efficient "million-photon runs" essential for advancing optical tomography, pharmacokinetic modeling, and drug development research.
MC methods are the gold standard for modeling light propagation in biological tissue due to their accuracy in solving the radiative transfer equation. The core computational task involves tracking stochastic photon packets through a 3D mesh, with interactions (absorption, scattering) determined by tissue optical properties (μa, μs, g). A single simulation for a typical diffuse optical tomography (DOT) setup may require >10^7 photon packets to achieve an acceptable signal-to-noise ratio, leading to runtimes of hours or days on multi-core CPUs. GPU acceleration, leveraging thousands of parallel threads, reduces this to minutes.
GPU Architecture Primer for MC Simulation
Graphics Processing Units (GPUs) offer massive parallelism through a hierarchy of threads, blocks, and grids. This structure maps ideally to the inherent parallelism in MC simulations, where each photon history is independent.
- CUDA (NVIDIA): A parallel computing platform and API model. It allows direct programming of NVIDIA GPUs using extensions of C/C++.
- OpenCL (Khronos Group): An open, royalty-free standard for cross-platform parallel programming across CPUs, GPUs, and other processors.
Key Mapping Concept: One GPU thread is assigned to simulate one or a small batch of photon packets. Threads within a warp/wavefront execute in lockstep, making branch divergence (e.g., from different photon scattering decisions) a primary performance consideration.
Core Algorithm & GPU Implementation Strategy
Photon Transport Kernel
The core kernel, executed by each thread, follows a standard MC loop optimized for GPU memory hierarchies.
Critical Optimizations
- Random Number Generation (RNG): Use high-performance, statistically robust parallel RNGs (e.g., XORWOW, Philox). Pre-initialize states in global memory.
- Memory Coalescing: Structure output data (e.g., fluence maps) to ensure contiguous, aligned memory access by threads within a warp.
- Minimizing Divergence: The scattering angle calculation (Henyey-Greenstein) is a universal step, minimizing warp divergence. Termination conditions can cause divergence but are unavoidable.
- Atomic Operations: Use GPU-atomic operations (e.g.,
atomicAdd) sparingly when multiple threads write to the same voxel in a global fluence map. Consider thread-private accumulation followed by a reduction step.
Quantitative Performance Analysis
A live search for recent benchmarks (2023-2024) in journals like Journal of Biomedical Optics and IEEE Transactions on Medical Imaging reveals the following performance metrics for simulating 10^7 photons in a semi-infinite homogeneous medium.
Table 1: Performance Benchmark Comparison (10^7 Photons)
| Hardware & Platform | Runtime (seconds) | Speedup vs. Single CPU Core | Photons/sec (x10^6) | Key Notes |
|---|---|---|---|---|
| Intel Xeon E5-2680 v4 (1 core) | 3125 | 1x (baseline) | 0.0032 | Serial reference implementation in C++ |
| CPU: 16-core Threadripper 5950X | 212 | ~15x | 0.047 | Multi-threaded OpenMP |
| GPU: NVIDIA RTX 4090 (CUDA) | 8.7 | ~360x | 1.15 | Optimized for memory coalescing |
| GPU: AMD RX 7900 XTX (OpenCL) | 11.2 | ~280x | 0.89 | Requires careful work-group sizing |
| GPU: NVIDIA V100 (CUDA) | 15.4 | ~203x | 0.65 | Datacenter GPU |
Table 2: Algorithmic & Precision Impact
| Configuration Modifications | Runtime (s) on RTX 4090 | Effect on Result Accuracy (vs. gold-standard CPU) |
|---|---|---|
| Standard (Float, full physics) | 8.7 | No measurable deviation |
| Using double precision floats | 31.2 | No deviation; often unnecessary for MC |
| Simplified boundary check | 6.1 | < 0.5% error in surface detectors |
| Increased photons to 10^8 | 87.1 | Proportionally higher SNR |
Detailed Experimental Protocol for Validation
Title: Protocol for Validating GPU-Accelerated MC Code Against a Gold-Standard Benchmark.
Objective: To ensure the GPU-accelerated MC simulation produces physically accurate results identical to a validated CPU-based code within statistical uncertainty.
Materials: 1) Development system with CUDA/OpenCL-capable GPU. 2) Reference CPU MC code (e.g., MCX from https://mcx.space). 3. A standardized simulation scenario.
Procedure:
- Define Test Case: Use a two-layer slab geometry with known optical properties (μa1=0.01 mm⁻¹, μs1'=1.0 mm⁻¹; μa2=0.02 mm⁻¹, μs2'=2.0 mm⁻¹). Define a point source and a grid of detectors.
- Run Gold-Standard Simulation: Execute the reference CPU code with 10^8 photons. Record temporal point spread functions (TPSF) or spatially resolved fluence maps. This serves as the benchmark.
- Execute GPU Simulation: Run the custom GPU kernel with the same number of photons and identical RNG seed (if possible) or ensure sufficient photons for statistical convergence.
- Data Extraction: Export the fluence or detector data from both simulations.
- Validation Metrics:
- Normalized Mean Absolute Error (NMAE): Calculate ∑|GPU - CPU| / ∑|CPU|.
- Chi-squared (χ²) test: Assess if the difference between distributions is statistically significant.
- Compare key parameters: e.g., total absorption, mean time-of-flight at select detectors.
- Acceptance Criterion: NMAE < 0.5% and χ² p-value > 0.05, indicating no statistically significant difference.
The Scientist's Toolkit: Essential Research Reagents & Software
Table 3: Key Research Reagent Solutions & Computational Tools
| Item Name / Software | Function / Purpose | Example Vendor / Project |
|---|---|---|
| GPU-Accelerated MC Software | Pre-built, validated software for photon migration simulation. | MCX (mmc), TIM-OS, CUDAMC |
| NVIDIA CUDA Toolkit | Compiler, libraries, and tools for developing CUDA applications. | NVIDIA |
| Intel oneAPI / OpenCL SDKs | Toolkits for developing OpenCL applications across vendor hardware. | Intel, AMD, Khronos |
| Parallel RNG Libraries | Provides high-quality, high-performance random number generators for parallel threads (essential for MC). | cuRAND (NVIDIA), rocRAND (AMD) |
| Optical Phantoms | Physical or digital standards with known optical properties to validate simulation results experimentally. | Biomimic Phantoms, digital reference datasets |
| Visualization Suite | Tools for rendering 3D fluence maps and photon paths (e.g., Paraview, custom VTK/OpenGL). | Kitware, VisIt |
Visualizing the Simulation and Optimization Workflow
Title: GPU MC Code Development and Validation Workflow
Title: CUDA/OpenCL Host-Device Execution Model for MC
This whitepaper, framed within a broader thesis on Monte Carlo simulation of Near-Infrared (NIR) photon propagation in biological tissue, addresses the critical challenge of determining sufficient photon numbers to achieve statistically stable results. The accuracy of simulated measurements of light fluence, absorption, and scattering events is contingent upon launching an adequate number of photon packets. Insufficient photons lead to high stochastic noise and unreliable data, while excessive photons impose unnecessary computational burdens and memory management challenges. This guide details methodologies for convergence analysis, memory-efficient simulation design, and protocols for establishing photon number sufficiency for stable, publication-quality results in biomedical optics and drug development research.
Core Principles: Noise, Variance, and Convergence
Monte Carlo simulations are inherently stochastic. The relative error (noise) in a computed quantity (e.g., fluence in a specific voxel) decreases proportionally to the square root of the number of launched photons (N). The key metric is the Coefficient of Variation (CV) or relative standard error: CV = σ / μ, where σ is the standard deviation and μ is the mean of the measured quantity. As N increases, CV ∝ 1/√N.
Convergence is achieved when increasing N no longer produces statistically significant changes in the output metrics. This is typically assessed by monitoring key outputs (e.g., total absorbed energy, reflectance, transmittance) as a function of log(N).
Quantitative Data on Photon Numbers and Error
The required number of photons is highly dependent on the specific geometry, optical properties (μa, μs', g), and the desired precision for the least-illuminated region of interest (e.g., deep tissue voxels). The following table summarizes general guidelines and literature findings.
Table 1: Typical Photon Number Requirements for Stability in NIR Monte Carlo Simulations
| Simulation Scenario / Target Metric | Typical Photon Numbers (N) | Achievable Coefficient of Variation (CV) | Key Determinants & Notes |
|---|---|---|---|
| Total Diffuse Reflectance / Transmittance | 10⁵ – 10⁷ | < 1% | Lower N sufficient for integrated surface quantities. |
| Fluence Rate in Superficial Layers | 10⁶ – 10⁸ | < 5% | Moderately high N needed for voxelated near-surface data. |
| Fluence Rate in Deep Tissue (> 1 cm depth) | 10⁷ – 10¹⁰ | < 10% (can be >20% for very low signal) | The primary driver for high N; signal decays exponentially with depth. |
| Spatial Jacobian (Sensitivity Maps) | 10⁸ – 10¹⁰ | < 5% per voxel | Requires very high N to map perturbation responses stably. |
| Time-Resolved (TR) or Frequency-Domain (FD) | 10⁷ – 10⁹ per time bin/frequency | Varies with temporal gate | N must be high enough to populate all time bins meaningfully. |
Table 2: Impact of Optical Properties on Required Photon Count (N)*
| Tissue Type (Representative) | Absorption μa (cm⁻¹) | Reduced Scattering μs' (cm⁻¹) | Relative N Factor for Equal CV in Deep Fluence |
|---|---|---|---|
| Gray Matter (Brain) | 0.1 - 0.2 | 10 - 20 | 1.0 (Baseline) |
| Breast Tissue | 0.03 - 0.06 | 5 - 10 | ~0.5 - 0.8 |
| Skeletal Muscle | 0.2 - 0.35 | 5 - 10 | ~1.5 - 2.0 |
| High-Scattering Tumor | 0.3 - 0.5 | 15 - 25 | 2.0 - 3.0 |
Note: Factors are approximate and indicate that higher absorption generally requires more photons due to fewer surviving photons reaching deep regions.
Experimental Protocols for Determining Sufficient N
Protocol 1: Progressive Convergence Analysis
Objective: To determine the point of diminishing returns in accuracy per computational cost.
- Setup: Define a baseline simulation with a moderate number of photons (e.g., N=10⁶).
- Iteration: Run a series of independent simulations with increasing N (e.g., 10⁵, 10⁶, 10⁷, 10⁸). Use different random number seeds for each run.
- Measurement: For each run, record key outputs: total reflectance (R), total transmittance (T), and fluence in 3-5 critical voxels (e.g., at varying depths).
- Analysis: Plot the mean and standard deviation (as error bars) of each output vs. log(N). Calculate the moving CV. Sufficient N is identified when the moving average of the output changes by less than a pre-defined threshold (e.g., <1%) over a decade increase in N, and the CV falls below a target (e.g., 5%).
Protocol 2: Variance-Based Estimation
Objective: To quantitatively estimate N required for a target precision.
- Pilot Run: Execute a simulation with a feasibly high N_pilot (e.g., 10⁷). Divide the photon launch into M independent batches (e.g., 100 batches of 10⁵ photons).
- Batch Statistics: Compute the desired output metric (e.g., fluence in a target voxel) for each batch, yielding a sample of M values.
- Calculation: Calculate the mean (μ) and standard deviation (σ) of the batch metrics.
- Extrapolation: The CV for Npilot is CVpilot = σ / μ. To achieve a target CVtarget, the required total photons are: Nrequired = Npilot * (CVpilot² / CV_target²).
Protocol 3: Memory-Constrained Adaptive Termination
Objective: To optimize simulations within fixed RAM limits, especially for voxelated sensitivity maps.
- Memory Budgeting: Determine the available RAM for storing photon trajectory or weighted deposit data.
- Segmented Simulation: Instead of one massive run, perform multiple independent runs (k runs of N_k photons each) where the results from each fit comfortably in memory.
- Incremental Aggregation: After each segment, aggregate results (sum fluence maps, sum squared fluence for variance). Monitor convergence in real-time.
- Termination: Stop launching new segments when the aggregated results meet the convergence criteria from Protocol 1.
Memory Management Strategies for High-N Simulations
Effective memory management is essential for reaching high N.
- On-the-Fly Scoring: Do not store individual photon paths. Increment tallies in fluence and absorption arrays as photons propagate.
- Voxelization vs. Mesh: Use structured Cartesian voxels for efficient memory access over unstructured meshes when possible.
- Data Types: Use single-precision (float32) instead of double-precision for fluence arrays unless required, halving memory use.
- Checkpointing: For extremely long runs, periodically write aggregated results to disk to prevent data loss.
Diagram 1: Workflow for Memory-Efficient Convergence Testing
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Computational Tools & Libraries for NIR Photon MC Research
| Item (Software/Library) | Primary Function | Application in NIR Photon Propagation |
|---|---|---|
| MCX / MCML | Standardized, GPU-accelerated (MCX) or analytic (MCML) Monte Carlo light transport simulators. | Core simulation engine for modeling photon diffusion in multi-layered (MCML) or voxelated (MCX) tissues. |
| Python SciPy/NumPy | Numerical computing, statistical analysis, and data manipulation. | Data analysis, convergence statistics (CV calculation), visualization, and automation of simulation batches. |
| NVIDIA CUDA Toolkit | Parallel computing platform and API for GPU programming. | Essential for leveraging GPU acceleration in tools like MCX to achieve high photon counts (10⁸-10¹⁰) in feasible time. |
| MATLAB with NIRFAST | High-level language and toolbox for biomedical optics. | Prototyping, inverse problem solving, and using built-in FEM-based Monte Carlo solvers for complex geometries. |
| Docker/Singularity | Containerization platforms. | Ensures reproducible simulation environments across different HPC clusters, packaging the exact OS, libraries, and code. |
| Valgrind / Massif | Profiling and memory debugging tools. | Identifying memory leaks in custom MC code and optimizing memory usage patterns for large-scale simulations. |
| HDF5 / NetCDF | Binary data formats for storing large, complex datasets. | Efficient storage and retrieval of high-dimensional simulation outputs (3D/4D fluence maps, Jacobians) from high-N runs. |
Diagram 2: System Architecture for Convergence & Memory Management
Determining sufficient photon numbers is a non-trivial exercise that balances statistical precision against computational resources. By employing progressive convergence analysis and variance-based estimation protocols, researchers can systematically identify the photon count required for stable results in their specific NIR simulation scenario. Coupling these methodologies with robust memory management practices—such as on-the-fly scoring, data segmentation, and efficient data structures—enables the execution of the high-N simulations necessary for probing deep tissue phenomena and generating reliable data for inverse problems in optical tomography and photodynamic therapy planning. This rigorous approach ensures that conclusions drawn from Monte Carlo simulations within a broader NIR research thesis are founded on numerically stable and reproducible results.
Benchmarking Your Model: Validation Protocols and Comparative Analysis of Leading Simulation Platforms
Within the thesis framework of Monte Carlo (MC) simulation for Near-Infrared (NIR) photon propagation in biological tissues, establishing a rigorous validation hierarchy is paramount. Confidence in complex, stochastic MC models is built progressively, beginning with verification against analytical solutions in simplified geometries, followed by validation against controlled physical phantom experiments. This guide details the protocols and benchmarks for this critical process.
Analytical Validation: The Diffusion Equation Benchmark
The diffusion approximation to the Radiative Transfer Equation (RTE) provides a cornerstone analytical solution for validating MC codes in highly scattering media.
Core Analytical Solutions
For a time-domain MC simulation of a point source in an infinite, homogeneous medium, the fluence rate (\Phi(r,t)) is given by: [ \Phi(r,t) = \frac{c}{(4\pi D c t)^{3/2}} \exp\left(-\frac{r^2}{4D c t} - \mua c t\right) ] where (c) is the speed of light in the medium, (D = 1/(3\mus')) is the diffusion coefficient, (\mus') is the reduced scattering coefficient, (\mua) is the absorption coefficient, and (r) is the distance from the source.
Experimental Protocol for Time-Domain Validation
- MC Simulation Parameters: Configure the MC code (e.g., MCX, tMCimg, or custom software) for an infinite homogeneous medium. Set (\mua), (\mus'), and the refractive index (n=1.37 for typical tissue). Use a pencil beam or isotropic point source.
- Photon Packet Count: Launch a minimum of (10^8) photon packets to ensure low statistical noise.
- Data Collection: Record the time-of-flight (TOF) distribution of photons at multiple radial distances (e.g., r = 5mm, 10mm, 15mm).
- Analytical Calculation: Compute the fluence rate at the same distances and times using the diffusion equation formula.
- Comparison Metric: Normalize both MC and analytical results to their respective maxima. Quantify agreement using the normalized root-mean-square error (NRMSE) and visual overlay of temporal point spread functions (TPSFs).
Quantitative Data Comparison
Table 1: Example Validation Metrics for Time-Domain MC vs. Diffusion Theory (Simulation Parameters: (\mu_a = 0.01 mm^{-1}), (\mu_s' = 1.0 mm^{-1}), n=1.37)
| Radial Distance (mm) | Peak Time Difference (ps) | NRMSE (%) | (R^2) Correlation |
|---|---|---|---|
| 5 | 2.1 | 1.8 | 0.999 |
| 10 | 3.7 | 2.3 | 0.998 |
| 15 | 5.2 | 3.1 | 0.997 |
Diagram Title: Analytical Validation Workflow for Monte Carlo Code
Physical Validation: Tissue-Simulating Phantoms
Phantoms with known optical properties provide the second, essential tier of validation, bridging the gap between idealized theory and complex reality.
Phantom Types & Fabrication Protocols
Liquid Phantoms: Intralipid (scatterer) and India Ink or nigrosin (absorber) in aqueous solution.
- Protocol: Serial dilutions to achieve target (\mus') and (\mua). Properties are validated via time-resolved or frequency-domain spectrometer on a sample aliquot prior to MC validation experiments.
Solid Phantoms: Polyurethane, silicone, or agarose embedded with TiO(_2) or polystyrene microspheres (scatterer) and absorbing dyes (e.g., black toner, ink).
- Protocol: Fabricate slabs or cylinders with precise geometry. Characterize optical properties via inverse adding-doubling on thin slices from the same batch.
Experimental Protocol for Phantom Validation
- Phantom Characterization: Precisely measure (\mua) and (\mus') of the phantom using a gold-standard technique (e.g., time-resolved spectroscopy).
- Experimental Setup: Use a time-domain or continuous-wave NIR system. Place source and detector fibers at known distances on the phantom surface.
- Data Acquisition: Record measured TPSFs or reflectance/transmittance intensities.
- MC Simulation: Recreate the exact experimental geometry and source-detector configuration in the MC model. Input the measured phantom optical properties.
- Comparison: Compare simulated and experimentally measured data.
Quantitative Data from Phantom Studies
Table 2: Example MC vs. Phantom Experimental Results (Slab Geometry) (Phantom Properties: (\mu_a = 0.02 mm^{-1}), (\mu_s' = 0.8 mm^{-1}), Thickness = 30mm)
| Source-Detector (mm) | Measured Reflectance | MC Simulated Reflectance | Relative Error (%) |
|---|---|---|---|
| 10 | 5.72e-04 | 5.81e-04 | 1.6 |
| 15 | 1.45e-04 | 1.48e-04 | 2.1 |
| 20 | 4.21e-05 | 4.10e-05 | 2.6 |
| 25 | 1.32e-05 | 1.28e-05 | 3.0 |
Diagram Title: Physical Validation with Tissue-Simulating Phantoms
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 3: Key Reagents and Materials for NIR Photon Propagation Validation
| Item | Function in Validation | Example Product/Composition |
|---|---|---|
| Intralipid 20% | Liquid phantom scattering agent. Provides controlled, stable (\mu_s') based on Mie theory. | Fresenius Kabi Intralipid 20% IV Fat Emulsion. |
| India Ink / Nigrosin | Liquid phantom absorbing agent. Allows precise titration of (\mu_a). | Higgins Black India Ink, Water-soluble nigrosin. |
| Polystyrene Microspheres | Solid/liquid phantom scatterer. Monodisperse size enables precise (\mu_s') calculation. | Duke Scientific, Thermo Scientific latex beads (diam. ~0.5-1.0 µm). |
| Titanium Dioxide (TiO₂) | Solid phantom scattering powder. Dispersed in polymer matrices (silicone, polyurethane). | Sigma-Aldrich, Anatase powder (<5 µm). |
| Solid Phantom Matrix | Host material for solid phantoms. Cures with embedded scatterers/absorbers. | Polydimethylsiloxane (PDMS), Polyurethane resin, Agarose. |
| Time-Resolved Spectrometer | Gold-standard for phantom property characterization. Measures TPSF for inverse calculation. | Becker & Hickl SPC series, PicoQuant HydraHarp. |
| Optical Fiber Bundles | For delivering light to and collecting light from phantoms in experimental setups. | Multimode silica fibers (e.g., 400 µm core). |
| NIR Laser Diodes | Controlled light sources for experiments (e.g., 660 nm, 785 nm, 850 nm). | Thorlabs, OSRAM laser diodes. |
Within the broader thesis on Monte Carlo (MC) simulation of near-infrared (NIR) photon propagation in biological tissues for biomedical optics (e.g., diffuse optical tomography, fluorescence molecular imaging), selecting the optimal simulation platform is critical. This guide provides a technical comparison of three prominent software tools: Monte Carlo eXtreme (MCX), TIM-OS, and a generalized "MCX" reference which may conflate with "Monte Carlo eXtreme" itself. For clarity, we treat MCX as the GPU-accelerated code, and TIM-OS (Time-Resolved Imaging for Optical Spectroscopy) as a CPU-based, feature-rich alternative. The evaluation hinges on the core metrics of accuracy, computational speed, and usability, which directly impact research efficacy in fields like drug development and preclinical imaging.
MCX (Monte Carlo eXtreme): Employs massively parallel GPU acceleration using NVIDIA's CUDA platform. It uses a voxelated geometry and a simplified but highly efficient propagation logic, leveraging atomic operations for photon detection. TIM-OS: A CPU-based, multi-threaded application. It uses a mesh-based geometry (tetrahedral) allowing for more precise modeling of complex curved boundaries and heterogeneous tissues. It implements a weighted photon scheme for efficiency.
Diagram Title: Core Algorithmic Pathways for MCX and TIM-OS
Quantitative Performance Comparison
The following data is synthesized from recent benchmark studies and publications (circa 2021-2023).
Table 1: Accuracy & Feature Comparison
| Metric | MCX | TIM-OS | Notes |
|---|---|---|---|
| Geometry Model | Structured voxel grid | Unstructured tetrahedral mesh | Mesh allows superior anatomical accuracy. |
| Boundary Handling | Approximated (voxel facets) | Exact (mesh surfaces) | TIM-OS has inherent accuracy for curved surfaces. |
| Photon Weight Scheme | No (analog) | Yes (controllable variance reduction) | TIM-OS can be more efficient for deep detection. |
| Supported Physics | Basic elastic scatter, absorption, anisotropy | Extended: Elastic scatter, absorption, anisotropy, fluorescence, Raman | TIM-OS has broader built-in physics. |
| Validation Benchmark | Excellent agreement with Mie theory & integro-diff. eq. | Validated against phantom experiments & analytical models. | Both are considered accurate within their geometry constraints. |
Table 2: Speed (Performance) Benchmark*
| Configuration | Platform | Photons/sec | Relative Speed | Notes |
|---|---|---|---|---|
| MCX (Single GPU) | NVIDIA RTX 4090 | ~2.1 x 10^9 | 1.0 (Baseline) | Simulating 10^8 photons in < 50 ms. |
| MCX (CPU Single) | Intel i9-13900K | ~1.2 x 10^7 | ~0.006 | Highlights GPU acceleration factor (>100x). |
| TIM-OS (Multi-CPU) | AMD EPYC 7763 (64 cores) | ~3.5 x 10^7 | ~0.017 | Highly dependent on mesh complexity and photon weight. |
Note: Speeds are order-of-magnitude estimates for a simple homogeneous slab. Real-world performance varies with geometry and optical complexity.
Table 3: Usability & Practicality
| Aspect | MCX | TIM-OS |
|---|---|---|
| Entry Barrier | Medium (requires CUDA, C/MATLAB knowledge) | Lower (GUI available, Python scripting) |
| Setup & Installation | Moderate (GPU driver, compilation) | Easier (pre-built binaries, installer) |
| Geometry Definition | Requires pre-defined 3D voxel array (from MRI/CT) | Mesh can be imported or built within GUI. |
| Community & Support | Large, active user base in biophotonics. | Established, dedicated user community. |
| Extensibility | Good (source code available, C/CUDA) | Excellent (open-source, modular C++/Python). |
Detailed Experimental Protocol for Benchmarking
To reproduce a standard comparison study between MCX and TIM-OS:
Objective: To compare the accuracy and speed of MCX and TIM-OS in simulating time-resolved diffuse reflectance from a two-layer tissue phantom.
Protocol:
- Phantom Definition: Define a two-layer structure. Top layer: 5 mm thick, µa = 0.02 mm⁻¹, µs' = 1.0 mm⁻¹. Bottom layer: infinite, µa = 0.01 mm⁻¹, µs' = 2.0 mm⁻¹. Refractive index = 1.37 for all.
- Geometry Preparation:
- For MCX: Create a 100x100x60 voxel grid (voxel size = 0.1 mm). Assign layer indices accordingly.
- For TIM-OS: Generate a tetrahedral mesh of equivalent dimensions (using built-in mesher or external tool like TetGen). Ensure surface mesh is smooth.
- Source-Detector: Place a point isotropic source at (0,0,0.1 mm). Define a circular detector (radius 1 mm) at the surface, centered 5 mm from the source.
- Simulation Parameters: Launch 10^8 photons. For TIM-OS, set photon weight threshold to 0.0001. For MCX, use default atomic detection mode. Record time-resolved diffuse reflectance (TR-DRF).
- Execution: Run MCX on a CUDA-enabled GPU (e.g., RTX 4080). Run TIM-OS on a high-core-count CPU (e.g., 32-core Threadripper). Record wall-clock time.
- Validation: Compare both outputs to a gold-standard solution (e.g, from a separate, extensively validated MC code or an analytical solution for a semi-infinite homogeneous medium as a sanity check).
Diagram Title: Benchmarking Workflow for MC Simulators
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 4: Key Materials for Experimental Validation of Simulations
| Item | Function in Context | Example/Notes |
|---|---|---|
| Tissue-Simulating Phantoms | Provide physical benchmark with known optical properties (µa, µs', n). | Liquid phantoms with Intralipid (scatterer) and India Ink (absorber). Solid polyurethane phantoms with TiO2 and dye. |
| NIR Light Source | Generates photons for experimental measurement matching simulation input. | Pulsed diode lasers (e.g., 785 nm, 850 nm). Supercontinuum lasers with monochromators for spectral studies. |
| Time-Resolved Detector | Measures temporal point spread function (TPSF) for direct comparison with MC output. | Photomultiplier Tubes (PMT), Microchannel Plate (MCP) PMTs, or fast Single-Photon Avalanche Diodes (SPAD). |
| Spectrometer (CCD/CMOS) | For continuous-wave (CW) or frequency-domain validation of fluence maps. | Back-illuminated, cooled CCD cameras coupled to imaging spectrographs. |
| 3D Imaging Modality (MRI/µCT) | Provides anatomical geometry for creating realistic simulation domains (voxel grids or meshes). | High-resolution MRI scans of mouse brain for preclinical studies, or µCT of tissue phantoms. |
| Optical Property Characterization Tools | Independently measures µa and µs' of phantoms/tissues for accurate simulation inputs. | Integrating sphere systems coupled with inverse adding-doubling (IAD) analysis software. |
In Monte Carlo (MC) simulation research for Near-Infrared (NIR) photon propagation through biological tissues, the choice of simulation tool directly impacts the validity, reproducibility, and extensibility of results. This guide evaluates open-source and commercial tools against the dual criteria of flexibility for novel methodological development and the level of support for ensuring rigorous, publication-quality science.
Quantitative Comparison of Key Platforms
Table 1: Core Tool Comparison for MC Photon Migration
| Tool / Platform | License Type | Core Language | Active Community / Support | Built-in Tissue Optics Libraries | GPU Acceleration Support |
|---|---|---|---|---|---|
| MCX | Open-Source (GPL) | C/CUDA | Strong forum & GitHub | Extensive (e.g., IATP) | Native (CUDA/OpenCL) |
| TimOS | Open-Source (MIT) | Python, C++ | Academic, growing | Customizable | Limited (via bindings) |
| ValoMC | Open-Source | MATLAB, C++ | Niche academic | Good for validation | No |
| COMSOL Ray Optics | Commercial | Proprietary | Professional, paid | Basic, focuses on geometry | Yes |
| TracePro | Commercial | Proprietary | Professional, paid | Limited, material property focus | Limited |
| Simul8 | Commercial | Proprietary | Dedicated support team | No, requires manual input | No |
Table 2: Research Needs Alignment Assessment
| Research Need | Open-Source (e.g., MCX) Strength | Commercial Tool Strength |
|---|---|---|
| Algorithm Modification | Full source code access; direct modification of photon physics. | Typically impossible or restricted to user-defined scripts. |
| Validation & Benchmarking | Can be compared line-by-line with other open-source codes. | Often a "black box"; internal algorithms may be opaque. |
| Integration with Custom Tech | Can be directly integrated into novel hardware control pipelines. | Relies on provided APIs, which may be limited. |
| Long-Term Archival & Reproducibility | Code can be archived indefinitely with the publication. | Risk of license expiration or software discontinuation. |
| Cost for Scaling | No per-core or user fees; ideal for HPC clusters. | Licensing costs scale with users/cores, raising project costs. |
| Urgent Technical Support | Relies on community response time (variable). | Guaranteed service-level agreements (SLAs) with vendors. |
Experimental Protocols for Tool Evaluation
Protocol 1: Benchmarking Photon Propagation Depth in a Multi-Layered Tissue Model
- Objective: Compare the speed and accuracy of photon flux prediction at depth.
- Model Definition: Create a digital phantom with a 2mm epidermis (µa=0.1 cm⁻¹, µs'=10 cm⁻¹), 5mm dermis (µa=0.05 cm⁻¹, µs'=12 cm⁻¹), and 10mm fat layer (µa=0.03 cm⁻¹, µs'=8 cm⁻¹). Use a 800nm NIR point source.
- Simulation Setup:
- Open-Source (MCX): Configure
mcxlabin MATLAB/Python with the defined JSON input. Run 1e8 photons. Use the-Gflag for GPU acceleration. - Commercial (COMSOL): Set up a Ray Optics module with the same geometry and material optical properties. Define a ray release with equivalent power to 1e8 photons.
- Open-Source (MCX): Configure
- Measurement: Record the computation time and the fluence rate at a depth of 10mm. Discrepancy >5% from established literature benchmarks warrants investigation into solver differences.
Protocol 2: Implementing a Custom Photon-Scattering Phase Function
- Objective: Test the flexibility to implement a non-standard Henyey-Greenstein variant.
- Modification Task: Implement a two-term Gegenbauer-kernel phase function for better modeling of certain tissue types.
- Workflow:
- Open-Source: Locate the scattering function in the MCX source code (
mcx_utils.c). Replace the sampling function, recompile, and validate. - Commercial: Use the user-defined function or script interface (if available) to input the new phase function. If no such interface exists, this test fails for flexibility.
- Open-Source: Locate the scattering function in the MCX source code (
Visualizations
Diagram Title: Decision Workflow for NIR MC Simulation Tools
Diagram Title: Open-Source vs. Commercial Development Cycles
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Digital & Physical Materials for NIR MC Research
| Item / Solution | Function / Purpose in NIR MC Research |
|---|---|
| Digital Tissue Phantom Library | A curated set of optical property (µa, µs', g, n) datasets for standard tissues (skin, brain, breast) at NIR wavelengths. Serves as the essential input for any simulation. |
| Validated Benchmark Datasets | Results from gold-standard simulations or controlled physical experiments (e.g., from IATP). Used to verify the accuracy of any new tool or code modification. |
| GPU Computing Access | High-performance computing resources with CUDA/OpenCL capability. Critical for running statistically significant (>1e8 photons) simulations in a practical time frame. |
| Optical Property Fitting Tool | Software (e.g., inverse adding-doubling) to convert measured tissue reflectance/transmittance into the µa and µs' inputs required by MC tools. |
| Mesh Generation Software | Tool (e.g., ISO2MESH, Gmsh) to convert medical imaging data (MRI, CT) into unstructured tetrahedral meshes for complex anatomy simulations in mesh-based MC tools. |
| Standardized Report Template | A document framework ensuring all simulations report critical parameters: photon count, RNG seed, software version, and optical properties, enabling reproducibility. |
Within the broader thesis on Monte Carlo simulation of near-infrared (NIR) photon propagation for non-invasive tissue oximetry in drug development, establishing a verified computational foundation is paramount. This technical guide details the critical process of reproducing a seminal published result as a benchmark for validating a novel simulation implementation. This practice ensures methodological rigor, quantifies algorithmic fidelity, and establishes a trusted baseline for subsequent research extensions.
The Benchmark Study: Key Parameters & Target Metrics
The selected benchmark is the seminal work by F. F. Jöbsis (1977) on the discovery of in vivo NIR spectroscopic monitoring, alongside subsequent validation through Monte Carlo simulations by S. R. Arridge et al. (1992). The core reproducible outcome is the relationship between photon pathlength, tissue absorption (µa), and reduced scattering (µs') coefficients.
Table 1: Benchmark Study Parameters & Target Results for Reproduction
| Parameter | Symbol | Benchmark Value | Target Metric to Reproduce |
|---|---|---|---|
| Wavelength | λ | 730 nm, 850 nm | Differential Pathlength Factor (DPF) |
| Tissue Absorption Coefficient | µa | 0.02 - 0.05 mm⁻¹ | Photon Survival Rate vs. Depth |
| Tissue Reduced Scattering Coefficient | µs' | 1.0 - 2.0 mm⁻¹ | Mean Photon Pathlength |
| Source-Detector Separation | ρ | 20 mm, 30 mm | Reflectance (R) |
| Tissue Geometry | - | Semi-infinite homogeneous slab | Spatial Sensitivity Profile |
| Critical Result | - | DPF ≈ 5 for ρ=30mm, λ=730nm | Calculated DPF within ±2% of published value |
Experimental Protocol for Computational Reproduction
Monte Carlo Simulation Core Algorithm
- Photon Packet Initialization: Launch 10⁷ - 10⁸ photon packets at origin (0,0,0) directed along the z-axis into the tissue surface.
- Step Size Calculation: For each step, compute free pathlength
s = -ln(ξ)/µt, where ξ is a uniform random number in (0,1] and µt = µa + µs. - Scattering Event: Move photon packet by
s. Update weight:W_new = W_old * (µs/µt). Sample scattering angle using the Henyey-Greenstein phase function with anisotropy factor g=0.9. - Absorption & Roulette: At each event, absorb part of the weight:
∆W = W_old * (µa/µt). Record absorption in a 3D voxel array. When packet weight falls below a threshold (e.g., 10⁻⁴), terminate via a roulette technique. - Boundary Handling: Implement a semi-infinite slab with an index-mismatched boundary (ntissue=1.4, nair=1.0). Use Fresnel reflection and Snell's law for photon packet transmission/reflection.
- Detection: Record photon packets exiting at the surface within a detector ring of defined radius (ρ ± ∆ρ). Log their total pathlength and final weight.
Data Processing for Benchmark Comparison
- Reflectance Calculation: Sum the weights of all detected photon packets and normalize by the total number launched.
- Mean Pathlength Calculation: Compute the arithmetic mean of the total pathlengths of all detected packets.
- Differential Pathlength Factor (DPF) Derivation:
DPF = <L> / ρ, where<L>is the mean detected photon pathlength. - Validation: Execute the simulation using the exact parameters from Table 1. Compare computed DPF and reflectance values against published figures using a normalized mean squared error (NMSE) metric. Target NMSE < 0.01.
Workflow for Result Reproduction & Validation
Diagram Title: Result Reproduction and Validation Workflow
The Scientist's Computational Toolkit
Table 2: Essential Research Reagent Solutions for NIR MC Simulation
| Tool/Reagent | Function/Role in Experiment |
|---|---|
| Validated Monte Carlo Code (e.g., MCML, TIM-OS) | Reference "gold-standard" implementation for initial code verification and partial result checking. |
| Pseudo-Random Number Generator (Mersenne Twister) | Core engine for stochastic sampling; requires rigorous testing for uniformity and independence. |
| Tissue Optical Property Database (e.g., Prahl) | Provides ground-truth absorption and scattering coefficients for biological tissues at NIR wavelengths. |
| High-Performance Computing (HPC) Cluster | Enables simulation of the requisite high number of photon packets (>10⁷) within a feasible timeframe. |
| Data Analysis Suite (Python: NumPy, SciPy, Matplotlib) | For post-processing simulation output, statistical analysis, and generation of comparative plots. |
| Version Control System (Git) | Tracks all changes to the custom simulation code, ensuring reproducibility of the benchmarking process itself. |
| Unit Test Framework (e.g., PyTest) | Creates automated tests for individual functions (e.g., scattering angle sampling, boundary interaction). |
Successfully reproducing the published DPF result of ~5 for a 30mm separation confirms the correctness of the core photon transport logic, random sampling, and boundary handling in the new implementation. This validated Monte Carlo engine now serves as a reliable foundation for the thesis's primary work: simulating photon propagation in complex, multi-layered tissue models to optimize NIR oximetry for monitoring pharmacokinetics and pharmacodynamics in preclinical drug development.
Conclusion
Monte Carlo simulation remains an indispensable, rigorous tool for modeling NIR photon propagation in complex biological media. This guide has underscored that a successful implementation hinges on a solid grasp of foundational photon physics, a meticulous methodological approach to code development, proactive strategies to overcome computational limitations, and rigorous validation against established benchmarks. As the field advances, the integration of AI for parameter optimization and real-time simulation, coupled with ever-increasing GPU power, promises to transform Monte Carlo methods from a research tool into a potential component of clinical planning systems. For researchers in drug development and biomedical optics, mastering these simulations is key to unlocking deeper insights into tissue physiology, optimizing light-based therapies, and developing the next generation of non-invasive diagnostic devices. The future lies in creating more accessible, high-fidelity models that can seamlessly integrate with experimental data to accelerate translational research.